Demecolcine
 | |
 | |
| Clinical data | |
|---|---|
| Other names | Colcemid |
| ATC code | |
| Identifiers | |
| |
| CAS Number | |
| PubChem CID | |
| ChemSpider | |
| UNII | |
| KEGG | |
| ChEMBL | |
| CompTox Dashboard (EPA) | |
| ECHA InfoCard | 100.006.832 |
| Chemical and physical data | |
| Formula | C21H25NO5 |
| Molar mass | 371.433 g·mol−1 |
| 3D model (JSmol) | |
| |
| |
| (verify) | |
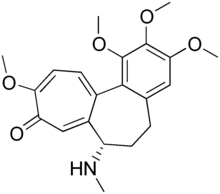
Demecolcine (INN; also known as colcemid) is a drug used in chemotherapy. It is closely related to the natural alkaloid colchicine with the replacement of the acetyl group on the amino moiety with methyl, but it is less toxic. It depolymerises microtubules and limits microtubule formation (inactivates spindle fibre formation), thus arresting cells in metaphase and allowing cell harvest and karyotyping to be performed.
During cell division, demecolcine inhibits mitosis at metaphase by inhibiting spindle formation. Medically, demecolcine has been used to improve the results of cancer radiotherapy by synchronising tumour cells at metaphase, the radiosensitive stage of the cell cycle.[1]
In animal cloning procedures, demecolcine makes an ovum eject its nucleus, creating space for insertion of a new nucleus.[2]
Mechanism of action
Demecolcine is a microtubule-depolymerizing drug like vinblastine. It acts by two distinct mechanisms. At very low concentration it binds to microtubule plus end to suppress microtubule dynamics.[3] Recent study has found at higher concentration demecolcine can promote microtubule detachment from microtubule organizing center. Detached microtubules with unprotected minus end depolymerize with time. Cytotoxicity of the cells seems to correlate better with microtubule detachment.[4] Lower concentration affects microtubule dynamics and cell migration.[4]
Research use
Demecolcine is used for scientific research in cells. It is used in a variety of ways, however, until recently, was used mostly for the study of mitosis in cells. For example, microtubules are necessary for the splitting of cells. More importantly, the movement of chromosomes during the M phase. Demecolcine inhibition of microtubules causes aneuploidy in mitotic cells where the microtubules fall apart or are suppressed before they can complete their function of pulling chromosomes into the daughter cell, also known as nondisjunction of chromosomes.[5] Demecolcine, depending on dose, has also been found to cause DNA fragmentation of chromosomes in micronuclei when nondisjunction occurs.[6]
References
- ^ Sutton M (February 1965). "Superior Mediastinal Obstruction Treated with Demecolcine Followed by Radiotherapy". British Medical Journal. 1 (5433): 495–6. doi:10.1136/bmj.1.5433.495. PMC 2165889. PMID 14238680.
- ^ Hou J, Lei T, Liu L, Cui X, An X, Chen Y (2006). "Demecolcine-induced enucleation of sheep meiotically maturing oocytes". Reproduction, Nutrition, Development. 46 (2): 219–26. doi:10.1051/rnd:2006002. PMID 16597428.
- ^ Jordan MA, Wilson L (April 2004). "Microtubules as a target for anticancer drugs". Nature Reviews. Cancer. 4 (4): 253–65. doi:10.1038/nrc1317. PMID 15057285. S2CID 10228718.
- ^ a b Yang H, Ganguly A, Cabral F (October 2010). "Inhibition of cell migration and cell division correlates with distinct effects of microtubule inhibiting drugs". The Journal of Biological Chemistry. 285 (42): 32242–50. doi:10.1074/jbc.M110.160820. PMC 2952225. PMID 20696757.
- ^ Hashimoto K, Todo T (July 2013). "Mitotic slippage underlies the relationship between p53 dysfunction and the induction of large micronuclei by colcemid". Mutagenesis. 28 (4): 457–64. doi:10.1093/mutage/get021. PMID 23702691.
- ^ Yamamoto M, Wakata A, Aoki Y, Miyamae Y, Kodama S (April 2014). "Chromosome loss caused by DNA fragmentation induced in main nuclei and micronuclei of human lymphoblastoid cells treated with colcemid". Mutation Research. 762: 10–6. doi:10.1016/j.mrfmmm.2014.02.002. PMID 24582839.