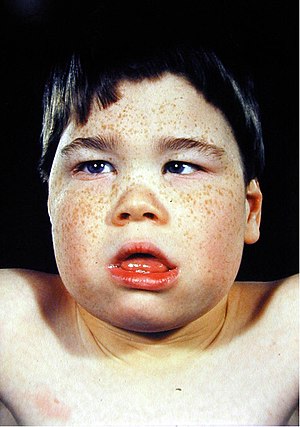
Alpha-thalassemia mental retardation syndrome
Alpha-thalassemia mental retardation syndrome (ATRX), also called alpha-thalassemia X-linked mental retardation, nondeletion type or ATR-X syndrome,[1] is an X-linked recessive condition associated with a mutation in the ATRX gene.[2] Males with this condition tend to be moderately intellectually disabled and have physical characteristics including coarse facial features, microcephaly (small head size), hypertelorism (widely spaced eyes), a depressed nasal bridge, a tented upper lip and an everted lower lip.[3] Mild or moderate anemia, associated with alpha-thalassemia, is part of the condition.[4] Females with this mutated gene have no specific signs or features, but if they do, they may demonstrate skewed X chromosome inactivation.
Symptoms and signs

The clinical presentation of this condition is as follows:[5]
- Delayed development
- Speech delayed
- Hypotonia
- Difficulty walking
- Pale skin (uncommon)
- Weakness (uncommon)
- Fatigue (uncommon)
Pathogenesis
"The role of ATRX as a regulator of heterochromatin dynamics raises the possibility that mutations in ATRX may lead to downstream transcriptional effects across the complex of genes or repetitive regions involved in the global context of the disorder, in addition to explaining phenotypical differences in these patients. For example, ATRX mutations affect the expression of alpha-globin gene cluster, causing alpha-thalassemia".[6] ATRX interacts with the transcription co-factor DAXX and the alpha-globin gene cluster.[6][7] Together they are all responsible for depositing the histone H3.3 at telomeric and pericentromeric regions. They are also responsible for regulating gene expression at these regions.[6] ATRX is characterized by hypo- and hypermethylated regions.[6] It's important to recognize that having a mutation in the ATRX gene does not necessarily guarantee that the patient has ATR-X syndrome.[6] However, it is common within ATR-X patients to have global hypermethylation of usually unmethylated regions, like CpG islands and promoters.[6] Several of the genes that undergo methylation changes are responsible for biosynthetic, metabolic, and methylation processes, and 42.5% of these genes are present in the telomeric and pericentromeric regions.[6] A couple of these genes include: PRDM9 and 2-BHMT2.[6] PRDM9 encodes for a histone H3 lysine-4 trimethyltransferase, which is a known target for ATRX, and 2-BHMT2 encodes for betaine-homocysteine methyltransferase, which catalyzes the methylation of homocysteine.[6]
ATR association can be separated into two groups. ATR-16 syndrome patients have a 1-2Mb deletion on the top of the chromosome 16 p-arm and are associated with a Mendelian inheritance of a-thalassemia.[8] ATR-X syndrome patients have no deletion in chromosome 16, a-thalassemia is rare, and this syndrome is consistent with X-linked recessive inheritance.[9] However, both groups have similar phenotypes.[10] The phenotypes resulting from ATR-X are due to skewed x-inactivation.[10] When X-inactivation occurs randomly, half of the cells in the carrier female would contain the abnormality.[10] When X-inactivation is skewed, more than 50% of one X chromosome are becoming inactive, and if that X-chromosome is passed to a male, they will have a higher percent of heterochromatin.[10] The ATR-X locus spans the control center Xist, which regulates X-inactivation.[11] When there is a XH2 mutation in the ATR-X locus, this indicates Xist to inactivate the chromosome increasing the amount of heterochromatin in males.[11]
Epigenetics is also present among transcriptional regulators. ATR-X is caused by XH2 mutations in the region Xq13.3, and XH2 belongs to the subgroup SNF2.[11] This group is important for regulating the transcription of the alpha genes.[11]
Diagnosis
The diagnosis of Alpha-thalassemia mental retardation syndrome can be done via genetic testing and clinical findings[12][13]
Treatment
Management for this condition is towards the mental disability and physical disabilities each case (individual) may present[12]
References
- ↑ Online Mendelian Inheritance in Man (OMIM): 301040
- ↑ Medina CF, Mazerolle C, Wang Y, et al. (March 2009). "Altered visual function and interneuron survival in Atrx knockout mice: inference for the human syndrome". Hum. Mol. Genet. 18 (5): 966–77. doi:10.1093/hmg/ddn424. PMID 19088125.
- ↑ Robert J. Gorlin; Meyer Michael Cohen; Raoul C. M. Hennekam (2001). Syndromes of the Head and Neck. Oxford University Press. p. 986. ISBN 0-19-511861-8.
- ↑ Stevenson, Roger E. (1993-01-01). Pagon, Roberta A.; Adam, Margaret P.; Ardinger, Holly H.; Wallace, Stephanie E.; Amemiya, Anne; Bean, Lora J.H.; Bird, Thomas D.; Dolan, Cynthia R.; Fong, Chin-To (eds.). Alpha-Thalassemia X-Linked Intellectual Disability Syndrome. Seattle (WA): University of Washington, Seattle. PMID 20301622. Archived from the original on 2021-01-25. Retrieved 2021-07-06.
- ↑ "Alpha thalassemia X-linked intellectual disability syndrome: MedlinePlus Genetics". medlineplus.gov. Archived from the original on 18 March 2021. Retrieved 15 August 2021.
- ↑ 6.0 6.1 6.2 6.3 6.4 6.5 6.6 6.7 6.8 (Schenkel et al., 2017)
- ↑ Iwase et al., 2011)
- ↑ (Higgs et al. 1989)
- ↑ (Harvey et al. 1990)
- ↑ 10.0 10.1 10.2 10.3 (Gibbons et al. 1992)
- ↑ 11.0 11.1 11.2 11.3 (Gibbons, Picketts, Villard, & Higgs, 1995)
- ↑ 12.0 12.1 RESERVED, INSERM US14-- ALL RIGHTS. "Orphanet: Alpha thalassemia X linked intellectual disability syndrome". www.orpha.net. Archived from the original on 28 February 2021. Retrieved 15 August 2021.
- ↑ Stevenson, Roger E. (1993). "Alpha-Thalassemia X-Linked Intellectual Disability Syndrome". GeneReviews®. University of Washington, Seattle. Archived from the original on 25 January 2021. Retrieved 15 August 2021.
- Schenkel, Laila; Kernoahn, Kristin; McBride, Arran; Reina, Ditta; Hodge, Amanda; Ainsworth, Peter; Rodenhiser, David; Pare, Guillaume; Berube, Nathalie; Skinner, Cindy; Boycott, Kym; Schwartz, Charles; Sadikovic, Bekim (2017). "Identification of epigenetic signature associated with alpha thalassemia/mental retardation X-linked syndrome". Epigenetics and Chromatin. 10 (1): 10. doi:10.1186/s13072-017-0118-4. PMC 5345252. PMID 28293299.
- Gibbons, R; Suthers, G; Wilkie, A; Buckle, V; Higgs, D (1992). "X-linked alpha-Thalassemia/Mental Retardation (ATR-X) Syndrome: Localization to Xq12-q21.31 by X inactivation and linkage analysis". American Journal of Human Genetics. 51 (5): 1136–1149. PMC 1682840. PMID 1415255.
- Gibbons, Richard; Picketts, David; Villard, Laurent; Higgs, Douglas (1995). "Mutations in a putative global transcriptional regulator cause X-linked mental retardation with alpha-Thalassemia (ATR-X Syndrome)". Cell. 80 (6): 837–845. doi:10.1016/0092-8674(95)90287-2. PMID 7697714. S2CID 16411046.
- Iwase, Shigeki; Xiang, Bin; Ghosh, Sharmistha; Ren, Ting; Lewis, Peter; Cochrane, Jesse; Allis, C; Picketts, David; Patel, Dinshaw; Li, Haitao; Shi, Yang (2011). "ATRX ADD domain links an atypical histone methylation recognition mechanism to human mental-retardation syndrome". Nature Structural and Molecular Biology. 18 (7): 769–776. doi:10.1038/nsmb.2062. PMC 3130887. PMID 21666679.
- Higgs, D; Vickers, M; Wilkie, A; Pretorius, I; Jarman, A; Weatherall, D (1989). "A review of the molecular genetics of the human alpha-globing gene cluster". Blood. 73 (5): 1081–1104. doi:10.1182/blood.V73.5.1081.1081. PMID 2649166.
- Harvey, M; Kearney, A; Smith, A; Trent, R (1990). "Occurrence of the alpha thalassaemie-mental retardation syndrome (non-deletional type) in an Australian male". Journal of Medical Genetics. 27 (9): 577–581. doi:10.1136/jmg.27.9.577. PMC 1017221. PMID 2231651.
Further reading
- GeneReviews/NCBI/NIH/UW entry on Alpha-Thalassemia X-Linked Mental Retardation Syndrome; ATRX Syndrome; Alpha Thalassemia/Mental Retardation, X-Linked; XLMR-Hypotonic Face Syndrome Archived 2021-01-25 at the Wayback Machine
- OMIM entries on Alpha-Thalassemia X-Linked Mental Retardation Syndrome Archived 2021-08-27 at the Wayback Machine
External links
| Classification |
|---|