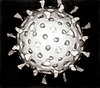
Chordopoxvirinae
| Chordopoxvirinae | |
|---|---|
| Virus classification | |
| (unranked): | Virus |
| Realm: | Varidnaviria |
| Kingdom: | Bamfordvirae |
| Phylum: | Nucleocytoviricota |
| Class: | Pokkesviricetes |
| Order: | Chitovirales |
| Family: | Poxviridae |
| Subfamily: | Chordopoxvirinae |
| Genera | |
Chordopoxvirinae is a subfamily of viruses in the family Poxviridae. Vertebrates and arthropods serve as natural hosts. Currently, 52 species are placed in this subfamily, divided among 18 genera. Diseases associated with this subfamily include smallpox.[1][2]
Four genera in this subfamily contain species that infect humans: Molluscipoxvirus, Orthopoxvirus, Parapoxvirus, and Yatapoxvirus.
Structure

The virions are generally enveloped though the intracellular mature virion form of the virus, which contains a different envelope and is also infectious. They vary in their shape depending upon the species but are generally shaped like a brick or as an oval form similar to a rounded brick because they are wrapped by the endoplasmic reticulum. The virion is exceptionally large, around 200 nm in diameter and 300 nm in length, and carries its genome in a single, double-stranded segment of DNA.[3]
Genomes are linear, around 130-375kb in length.[1]
| Genus | Structure | Symmetry | Capsid | Genomic arrangement | Genomic segmentation |
|---|---|---|---|---|---|
| Yatapoxvirus | Brick-shaped | Enveloped | Linear | Monopartite | |
| Cervidpoxvirus | Brick-shaped | Enveloped | Linear | Monopartite | |
| Leporipoxvirus | Brick-shaped | Enveloped | Linear | Monopartite | |
| Suipoxvirus | Brick-shaped | Enveloped | Linear | Monopartite | |
| Molluscipoxvirus | Brick-shaped | Enveloped | Linear | Monopartite | |
| Crocodylidpoxvirus | Ovoid | Enveloped | Linear | Monopartite | |
| Capripoxvirus | Brick-shaped | Enveloped | Linear | Monopartite | |
| Orthopoxvirus | Brick-shaped | Enveloped | Linear | Monopartite | |
| Avipoxvirus | Brick-shaped | Enveloped | Linear | Monopartite | |
| Parapoxvirus | Ovoid | Enveloped | Linear | Monopartite |
Lifecycle
Viral replication is cytoplasmic. Entry into the host cell is achieved by attachment of the viral proteins to host glycosaminoglycans (GAGs), which mediate endocytosis of the virus into the host cell. Fusion with the plasma membrane releases the core into the host cytoplasm. In the early phase, early genes are transcribed in the cytoplasm by viral RNA polymerase. Early expression begins at 30 minutes postinfection. The core is completely uncoated as early expression ends, the viral genome is then free in the cytoplasm. In the intermediate phase, intermediate genes are expressed, triggering genomic DNA replication about 100 minutes after infection. In the late phase, late genes are expressed from 140 min to 48 hours postinfection, producing all structural proteins. Assembly of progeny virions starts in cytoplasmic viral factories, producing a spherical, immature particle. This virus particle matures into brick-shaped intracellular mature virion, which can be released upon cell lysis, or can acquire a second double membrane from trans-Golgi and bud as external enveloped virion host receptors, which mediates endocytosis. Replication follows the DNA strand displacement model. DNA-templated transcription is the method of transcription. The virus exits the host cell by microtubular outwards viral transport, and exists in occlusion bodies after cell death and remains infectious until finding another host. Humans, vertebrates, and arthropods serve as the natural host. Transmission routes are fomite, contact, and airborne particles.[1]
| Genus | Host details | Tissue tropism | Entry details | Release details | Replication site | Assembly site | Transmission |
|---|---|---|---|---|---|---|---|
| Yatapoxvirus | Monkeys; baboons | None | Glycosaminoglycans | Lysis; budding | Cytoplasm | Cytoplasm | Contact; insects |
| Cervidpoxvirus | Deer | None | Glycosaminoglycans | Lysis; budding | Cytoplasm | Cytoplasm | Unknown |
| Leporipoxvirus | Lagomorph; squirrels | None | Glycosaminoglycans | Lysis; budding | Cytoplasm | Cytoplasm | Arthropods; contact |
| Suipoxvirus | Swine | None | Glycosaminoglycans | Lysis; budding | Cytoplasm | Cytoplasm | Unknown |
| Molluscipoxvirus | Humans | None | Glycosaminoglycans | Lysis; budding | Cytoplasm | Cytoplasm | Sex; contact |
| Crocodylidpoxvirus | Crocodiles | None | Glycosaminoglycans | Lysis; budding | Cytoplasm | Cytoplasm | Unknown |
| Capripoxvirus | Sheep; goat; cattle | None | Glycosaminoglycans | Lysis; budding | Cytoplasm | Cytoplasm | Arthropods; contact |
| Orthopoxvirus | Humans; mammals | None | Glycosaminoglycans | Lysis; budding | Cytoplasm | Cytoplasm | Variola virus; Respiratory; contact; zoonosis |
| Avipoxvirus | Birds | None | Glycosaminoglycans | Lysis; budding | Cytoplasm | Cytoplasm | Arthropods; aerosol |
| Parapoxvirus | Humans; mammals | None | Glycosaminoglycans | Lysis; budding | Cytoplasm | Cytoplasm | Zoonosis; contact |
Taxonomy
The classification in this subfamily is based on the morphology, nucleic acid type, mode of replication, host organisms, and type of disease caused.
Nine genera in this subfamily are recognized; also, a number of species have not yet been assigned to a genus.
The species in the genus Avipoxvirus infect birds; those in the genera Caiman poxvirus and Crocodylipoxvirus both infect crocodilians.
The other genera in this subfamily infect mammals.
The following genera are recognized:[2]
Evolution
The last common ancestor of the extant poxviruses that infect vertebrates existed 0.5 million years ago. The genus Avipoxvirus diverged from the ancestor 249 ± 69 thousand years ago. The ancestor of the genus Orthopoxvirus was next to diverge from the other clades at 0.3 million years ago. A second estimate of this divergence time places this event at 166 ± 43,000 years ago.[4] The division of the Orthopox into the extant species occurred about 14,000 years ago. The genus Leporipoxvirus diverged around 137 ± 35,000 years ago. This was followed by the ancestor of the genus Yatapoxvirus. The last common ancestor of the Capripoxvirus and Suipoxvirus diverged 111 ± 29,000 years ago.[citation needed]
A Bayesian study of orthopox genomes suggests that the unclassified Yoka poxvirus diverged from the lineage that gave rise to the orthopoxviruses roughly 90,000 years ago.[5] The orthopox viruses diverged from the other pox viruses about 10,000 years ago. Camelpox, taterapox, and variola viruses arose 3,500 years ago and horsepox virus 3,000 years ago. These viruses may have arisen in the Horn of Africa.[citation needed]
Another Bayesian study suggests that variola arose about 3500 years ago.[6]
References
- ↑ 1.0 1.1 1.2 1.3 "Viral Zone". ExPASy. Archived from the original on 19 June 2015. Retrieved 15 June 2015.
- ↑ 2.0 2.1 "Virus Taxonomy: 2019 Release". talk.ictvonline.org. International Committee on Taxonomy of Viruses. Archived from the original on 20 March 2020. Retrieved 9 May 2020.
- ↑ International Committee on Taxonomy of Viruses (15 June 2004). "ICTVdb Descriptions: 58. Poxviridae". Archived from the original on 3 July 2007. Retrieved 26 February 2005.
- ↑ Babkin IV, Babkina IN (2011) Molecular dating in the evolution of vertebrate poxviruses. Intervirology 54(5):253-260. doi:10.1159/000320964.
- ↑ Babkin IV, Babkina IN (2012) A retrospective study of the orthopoxvirus molecular evolution. Infect Genet Evol
- ↑ Babkin IV, Shelkunov SN (2008) Molecular evolution of poxviruses. Genetika 44(8):1029-1044
External links
- Viralzone: Chordopoxvirinae Archived 5 January 2017 at the Wayback Machine
- ICTV Archived 10 July 2015 at the Wayback Machine
- Electron micrographs - [1] Archived 18 February 2009 at the Wayback Machine