Ribose-5-phosphate isomerase
| ribose-5-phosphate isomerase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
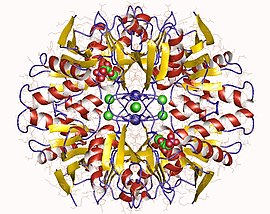 D-Ribose-5-phosphate isomerase homotetramer, Pyrococcus horikoshii | |||||||||
| Identifiers | |||||||||
| EC no. | 5.3.1.6 | ||||||||
| CAS no. | 9023-83-0 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
Ribose-5-phosphate isomerase (Rpi) encoded by the RPIA gene is an enzyme (EC 5.3.1.6) that catalyzes the conversion between ribose-5-phosphate (R5P) and ribulose-5-phosphate (Ru5P). It is a member of a larger class of isomerases which catalyze the interconversion of chemical isomers (in this case structural isomers of pentose). It plays a vital role in biochemical metabolism in both the pentose phosphate pathway and the Calvin cycle. The systematic name of this enzyme class is D-ribose-5-phosphate aldose-ketose-isomerase.
Structure
Gene
RpiA in human beings is encoded on the second chromosome on the short arm (p arm) at position 11.2. Its encoding sequence is nearly 60,000 base pairs long.[1] The only known naturally occurring genetic mutation results in ribose-5-phosphate isomerase deficiency, discussed below. The enzyme is thought to have been present for most of evolutionary history. Knock-out experiments conducted on the genes of various species meant to encode RpiA have indicated similar conserved residues and structural motifs, indicating ancient origins of the gene. [2]
Protein

Rpi exists as two distinct proteins, termed RpiA and RpiB. Although RpiA and RpiB catalyze the same reaction, they show no sequence or overall structural homology. According to Jung et al.,[3] an assessment of RpiA using SDS-PAGE shows that the enzyme is a homodimer of 25 kDa subunits. The molecular mass of the RpiA dimer was found to be 49 kDa [3] by gel filtration. Recently, the crystal structure of RpiA was determined. (please see http://www3.interscience.wiley.com/cgi-bin/fulltext/97516673/PDFSTAR[permanent dead link])
Due to its role in the pentose phosphate pathway and the Calvin cycle, RpiA is highly conserved in most organisms, such as bacteria, plants, and animals. RpiA plays an essential role in the metabolism of plants and animals, as it is involved in the Calvin cycle which takes place in plants, and the pentose phosphate pathway which takes place in plants as well as animals.
All orthologs of the enzyme maintain an asymmetric tetramer quaternary structure with a cleft containing the active site. Each subunit consists of a five stranded β-sheet. These β-sheets are surrounded on both sides by α-helices.[4] This αβα motif is not uncommon in other proteins, suggesting possible homology with other enzymes.[5] The separate molecules of the enzyme are held together by highly polar contacts on the external surfaces of the monomers. It is presumed that the active site is located where multiple β-sheet C termini come together in the enzymatic cleft. This cleft is capable of closing upon recognition of the phosphate on the pentose (or an appropriate phosphate inhibitor). The active site is known to contain conserved residues equivalent to the E. coli residues Asp81, Asp84, and Lys94. These are directly involved in catalysis.[6]
Mechanism
In the reaction, the overall consequence is the movement of a carbonyl group from carbon number 1 to carbon number 2; this is achieved by the reaction going through an enediol intermediate (Figure 1).[6] Through site-directed mutagenesis, Asp87 of spinach RpiA was suggested to play the role of a general base in the interconversion of R5P to Ru5P.[7]
The first step in the catalysis is the docking of the pentose into the active site in the enzymatic cleft, followed by allosteric closing of the cleft. The enzyme is capable of bonding with the open-chain or ring form of the sugar-phosphate. If it does bind the furanose ring, it next opens the ring. Then the enzyme forms the eneldiol which is stabilized by a lysine or arginine residue.[6][8] Calculations have demonstrated that this stabilization is the most significant contributor to the overall catalytic activity of this isomerase and a number of others like it.[9]
Function
The protein encoded by RPIA gene is an enzyme, which catalyzes the reversible conversion between ribose-5-phosphate and ribulose-5-phosphate in the pentose-phosphate pathway. This gene is highly conserved in most organisms. The enzyme plays an essential role in the carbohydrate metabolism. Mutations in this gene cause ribose 5-phosphate isomerase deficiency. A pseudogene is found on chromosome 18.[10]
Pentose phosphate pathway
In the non-oxidative part of the pentose phosphate pathway, RPIA converts Ru5P to R5P which then is converted by ribulose-phosphate 3-epimerase to xylulose-5-phosphate (figure 3).[11] The result of the reaction essentially is the conversion of the pentose phosphates to intermediates used in the glycolytic pathway. In the oxidative part of the pentose phosphate pathway, RpiA converts Ru5P to the final product, R5P through the isomerization reaction (figure 3). The oxidative branch of the pathway is a major source for NADPH which is needed for biosynthetic reactions and protection against reactive oxygen species.[12]
Calvin cycle
In the Calvin cycle, the energy from the electron carriers is used in carbon fixation, the conversion of carbon dioxide and water into carbohydrates. RPIA is essential in the cycle, as Ru5P generated from R5P is subsequently converted to ribulose-1,5-bisphosphate (RuBP), the acceptor of carbon dioxide in the first dark reaction of photosynthesis (Figure 3).[13] The direct product of RuBP carboxylase reaction is glyceraldehyde-3-phosphate; these are subsequently used to make larger carbohydrates.[14] Glyceraldehyde-3-phosphate is converted to glucose which is later converted by the plant to storage forms (e.g., starch or cellulose) or used for energy.[15]
Clinical significance
Ribose-5-phosphate isomerase deficiency is mutated in a rare disorder, Ribose-5-phosphate isomerase deficiency. The disease has only one known affected patient, diagnosed in 1999.[16] It has been found to be caused by a combination of two mutations. The first is an insertion of a premature stop codon into the gene encoding the isomerase, and the second is a missense mutation. The molecular pathology is, as yet, unclear.[17]
RpiA and hepatocarcinogenesis
Human ribose-5-phosphate isomerase A (RpiA) plays a role in human hepatocellular carcinoma (HCC).[18] A significant increase in RpiA expression was detected both in tumor biopsies of HCC patients and in a liver cancer tissue array. Importantly, the clinicopathological analysis indicated that RpiA mRNA levels were highly correlated with clinical stage, grade, tumor size, types, invasion and alpha-fetoprotein levels in the HCC patients. In addition, the ability of RpiA to regulate cell proliferation and colony formation in different liver cancer cell lines required ERK signaling as well as the negative modulation of PP2A activity and that the effects of RpiA could be modulated by the addition of either a PP2A inhibitor or activator. It suggests that RpiA overexpression can induce oncogenesis in HCC.[19]
RpiA and the malaria parasite
RpiA generated attention when the enzyme was found to play an essential role in the pathogenesis of the parasite Plasmodium falciparum, the causative agent of malaria. Plasmodium cells have a critical need for a large supply of the reducing power of NADPH via PPP in order to support their rapid growth. The need for NADPH is also required to detoxify heme, the product of hemoglobin degradation.[20] Furthermore, Plasmodium has an intense requirement for nucleic acid production to support its rapid proliferation. The R5P produced via increased pentose phosphate pathway activity is used to generate 5-phospho-D-ribose α-1-pyrophosphate (PRPP) needed for nucleic acid synthesis. It has been shown that PRPP concentrations are increased 56 fold in infected erythrocytes compared with uninfected erythrocytes.[17] Hence, designing drugs that target RpiA in Plasmodium falciparum could have therapeutic potential for patients that suffer from malaria.
Interactions
RPIA has been shown to interact with PP2A.[19]
Structural studies
As of late 2007, 15 structures have been solved for this class of enzymes, with PDB accession codes 1LK5, 1LK7, 1LKZ, 1M0S, 1NN4, 1O1X, 1O8B, 1UJ4, 1UJ5, 1UJ6, 1USL, 1XTZ, 2BES, 2BET, and 2F8M.
References
- ^ U.S. National Library of Medicine http://ghr.nlm.nih.gov/gene/RPIA
- ^ Sørensen KI, Hove-Jensen B (February 1996). "Ribose catabolism of Escherichia coli: characterization of the rpiB gene encoding ribose phosphate isomerase B and of the rpiR gene, which is involved in regulation of rpiB expression". Journal of Bacteriology. 178 (4): 1003–11. doi:10.1128/jb.178.4.1003-1011.1996. PMC 177759. PMID 8576032.
- ^ a b Jung CH, Hartman FC, Lu TY, Larimer FW (January 2000). "D-ribose-5-phosphate isomerase from spinach: heterologous overexpression, purification, characterization, and site-directed mutagenesis of the recombinant enzyme". Archives of Biochemistry and Biophysics. 373 (2): 409–17. doi:10.1006/abbi.1999.1554. PMID 10620366. S2CID 13217828.
- ^ Zhang RG, Andersson CE, Skarina T, Evdokimova E, Edwards AM, Joachimiak A, Savchenko A, Mowbray SL (October 2003). "The 2.2 Å resolution structure of RpiB/AlsB from Escherichia coli illustrates a new approach to the ribose-5-phosphate isomerase reaction". Journal of Molecular Biology. 332 (5): 1083–94. doi:10.1016/j.jmb.2003.08.009. PMC 2792017. PMID 14499611.
- ^ Rossmann MG, Moras D, Olsen KW (July 1974). "Chemical and biological evolution of nucleotide-binding protein". Nature. 250 (463): 194–9. Bibcode:1974Natur.250..194R. doi:10.1038/250194a0. PMID 4368490. S2CID 4273028.
- ^ a b c Zhang Rg, Andersson CE, Savchenko A, Skarina T, Evdokimova E, Beasley S, Arrowsmith CH, Edwards AM, Joachimiak A, Mowbray SL (January 2003). "Structure of Escherichia coli ribose-5-phosphate isomerase: a ubiquitous enzyme of the pentose phosphate pathway and the Calvin cycle". Structure. 11 (1): 31–42. doi:10.1016/S0969-2126(02)00933-4. PMC 2792023. PMID 12517338.
- ^ Gengenbacher M, Fitzpatrick TB, Raschle T, Flicker K, Sinning I, Müller S, Macheroux P, Tews I, Kappes B (February 2006). "Vitamin B6 biosynthesis by the malaria parasite Plasmodium falciparum: biochemical and structural insights". The Journal of Biological Chemistry. 281 (6): 3633–41. doi:10.1074/jbc.M508696200. PMID 16339145.
- ^ Woodruff, William W.; Wolfenden, Richard (18 July 1978). "Inhibition of ribose-5-phosphate by 4-phosphoerythronate". Journal of Biological Chemistry. 254 (13). Retrieved 6 March 2013.
- ^ Feierberg I, Åqvist (2002). "Computational modeling of enzymatic keto–enol isomerization reactions". Theoretical Chemistry Accounts. 108 (2): 71–84. doi:10.1007/s00214-002-0365-7. S2CID 95097042.
- ^ "Entrez Gene: RPIA ribose 5-phosphate isomerase A".
- ^ Berg, Jeremy M. (2012). Biochemistry. New York: W.H. Freeman and Company. ISBN 978-1-4292-2936-4.
- ^ Struzyńska L, Chalimoniuk M, Sulkowski G (September 2005). "The role of astroglia in Pb-exposed adult rat brain with respect to glutamate toxicity". Toxicology. 212 (2–3): 185–94. doi:10.1016/j.tox.2005.04.013. PMID 15955607.
- ^ Martin W, Henze K, Kellerman J, Flechner A, Schnarrenberger C (February 1996). "Microsequencing and cDNA cloning of the Calvin cycle/OPPP enzyme ribose-5-phosphate isomerase (EC 5.3.1.6) from spinach chloroplasts". Plant Molecular Biology. 30 (4): 795–805. doi:10.1007/BF00019012. PMID 8624410. S2CID 4218453.
- ^ A. A. Benson; J. A. Bassham; M. Calvin; T. C. Goodale; V. A. Haas; W. Stepka (1950). "The Path of Carbon in Photosynthesis. V. Paper Chromatography and Radioautography of the Products1". Journal of the American Chemical Society. 72 (4): 1710–1718. doi:10.1021/ja01160a080.
- ^ Nelson, David L. (2005). Principles of Biochemistry. New Yord: W.H Freeman and Company. ISBN 0-7167-4339-6.
- ^ Wamelink MM, Grüning NM, Jansen EE, Bluemlein K, Lehrach H, Jakobs C, Ralser M (September 2010). "The difference between rare and exceptionally rare: molecular characterization of ribose 5-phosphate isomerase deficiency" (PDF). Journal of Molecular Medicine. 88 (9): 931–9. doi:10.1007/s00109-010-0634-1. hdl:1871/34686. PMID 20499043. S2CID 10870492.
- ^ a b Huck JH, Verhoeven NM, Struys EA, Salomons GS, Jakobs C, van der Knaap MS (April 2004). "Ribose-5-phosphate isomerase deficiency: new inborn error in the pentose phosphate pathway associated with a slowly progressive leukoencephalopathy". American Journal of Human Genetics. 74 (4): 745–51. doi:10.1086/383204. PMC 1181951. PMID 14988808.
- ^ Riganti C, Gazzano E, Polimeni M, Aldieri E, Ghigo D (August 2012). "The pentose phosphate pathway: an antioxidant defense and a crossroad in tumor cell fate". Free Radical Biology & Medicine. 53 (3): 421–36. doi:10.1016/j.freeradbiomed.2012.05.006. PMID 22580150.
- ^ a b Ciou SC, Chou YT, Liu YL, Nieh YC, Lu JW, Huang SF, Chou YT, Cheng LH, Lo JF, Chen MJ, Yang MC, Yuh CH, Wang HD (July 2015). "Ribose-5-phosphate isomerase A regulates hepatocarcinogenesis via PP2A and ERK signaling". International Journal of Cancer. 137 (1): 104–15. doi:10.1002/ijc.29361. PMID 25429733.
- ^ Becker K, Rahlfs S, Nickel C, Schirmer RH (April 2003). "Glutathione--functions and metabolism in the malarial parasite Plasmodium falciparum". Biological Chemistry. 384 (4): 551–66. doi:10.1515/BC.2003.063. PMID 12751785. S2CID 20083367.
- Dickens F, Williamson DH (November 1956). "Pentose phosphate isomerase and epimerase from animal tissues". The Biochemical Journal. 64 (3): 567–78. doi:10.1042/bj0640567. PMC 1199776. PMID 13373810.
- Horecker BL, Smyrniotis PZ, Seegmiller JE (November 1951). "The enzymatic conversion of 6-phosphogluconate to ribulose-5-phosphate and ribose-5-phosphate". The Journal of Biological Chemistry. 193 (1): 383–96. doi:10.1016/S0021-9258(19)52464-4. PMID 14907726.
- Hurwitz J, Weissbach A, Horecker BL, Smyrniotis PZ (February 1956). "Spinach phosphoribulokinase". The Journal of Biological Chemistry. 218 (2): 769–83. doi:10.1016/S0021-9258(18)65841-7. PMID 13295229.
