Orthohantavirus
| Orthohantavirus | |
|---|---|

| |
| Transmission electron micrograph of Sin Nombre orthohantavirus | |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Negarnaviricota |
| Class: | Ellioviricetes |
| Order: | Bunyavirales |
| Family: | Hantaviridae |
| Subfamily: | Mammantavirinae |
| Genus: | Orthohantavirus |
| Type species | |
| Hantaan orthohantavirus | |
| Synonyms[2] | |
|
Hantavirus | |
Orthohantavirus is a genus of single-stranded, enveloped, negative-sense RNA viruses in the family Hantaviridae of the order Bunyavirales.[3] Members of this genus may be called orthohantaviruses or simply hantaviruses. They normally cause infection in rodents, but do not cause disease in them.[3] Humans may become infected with hantaviruses through contact with rodent urine, saliva, or feces. Some strains cause potentially fatal diseases in humans, such as hantavirus hemorrhagic fever with renal syndrome (HFRS), or hantavirus pulmonary syndrome (HPS), also known as hantavirus cardiopulmonary syndrome (HCPS),[4] while others have not been associated with known human disease.[5] HPS (HCPS) is a "rare respiratory illness associated with the inhalation of aerosolized rodent excreta (urine and feces) contaminated by hantavirus particles."[4]
Characteristics
Structure
Hantavirus virions are about 120–160 nanometers (nm) in diameter. The lipid bilayer of the viral envelope is about 5 nm thick and is embedded with viral surface proteins to which sugar residues are attached. These glycoproteins, known as Gn and Gc, are encoded by the M segment of the viral genome. They tend to associate (heterodimerize) with each other and have both an interior tail and an exterior domain that extends to about 6 nm beyond the envelope surface.[citation needed]
Inside the envelope are the nucleocapsids. These are composed of many copies of the nucleocapsid protein N, which interact with the three segments of the viral genome to form helical structures. The virally encoded RNA polymerase is also found in the interior. By mass, the virion is greater than 50% protein, 20–30% lipid and 2–7% carbohydrate. The density of the virions is 1.18 gram per cubic centimeter. These features are common to all members of the family Hantaviridae.[citation needed]
Genome
The genome of hantaviruses is negative-sense, single-stranded RNA. Their genomes are composed of three segments: the small (S), medium (M), and large (L) segments. The S segment, 1-3 kilobases (kb) in length, encodes for the nucleocapsid (N) protein. The M segment, 3.2-4.9 kb in length, encodes a glycoprotein precursor polyprotein that is co-translationally cleaved into the envelope glycoproteins Gn and Gc, alternatively called G1 and G2. The L segment, 6.8–12 kb in length, encodes the L protein which functions primarily as the viral RNA-dependent RNA polymerase used for transcription and replication.[6][7]
Within virions, the genomic RNAs of hantaviruses are thought to complex with the N protein to form helical nucleocapsids, the RNA component of which circularizes due to sequence complementarity between the 5' and 3' terminal sequences of genomic segments.[citation needed]
As with other Bunyavirales, each of the three segments has a consensus 3'-terminal nucleotide sequence (AUCAUCAUC), which is complementary to the 5'-terminal sequence and is distinct from those of the other four genera in the family.[8] These sequences appear to form panhandle structure which seem likely to play a role in replication and encapsidation facilitated by binding with the viral nucleocapsid (N) protein.[9] The large segment is 6530–6550 nucleotides (nt) in length, the medium is 3613–3707 nt in length and the small is 1696–2083 nt in length.[citation needed]
No nonstructural proteins are known, unlike the other genera in this family. At the 5' and 3' of each segment are short noncoding sequences: the noncoding segment in all sequences at the 5' end is 37–51 nt. The 3' noncoding regions differ: L segment 38–43 nt; M segment 168–229 nt; and S segment 370–730 nt. The 3' end of the S segment is conserved between the genera suggesting a functional role.[citation needed]
Life cycle
Viral entry into host cells initiates by binding to surface cell receptors. Integrins are considered to be the main receptors for hantaviruses in vitro but complement decay-accelerating factor (DAF) and globular heads of complement C1q receptor (gC1qR) have mediated attachment in cultured cells too. Entry may proceed through a number of possible routes, including clathrin-dependent endocytosis, clathrin-independent receptor-mediated endocytosis, and micropinocytosis. Viral particles are then transported to late endosomes. Gc-mediated membrane fusion with the endosomal membrane, triggered by low pH, releases the nucleocapsid into the cytoplasm.[7]
After the release of the nucleocapsids into cytoplasm, the complexes are targeted to the ER–Golgi Intermediate compartments (ERGIC) through microtubular-associated movement resulting in the formation of viral factories at ERGIC.[citation needed]
These factories then facilitate transcription and subsequent translation of the viral proteins. Transcription of viral genes must be initiated by association of the L protein with the three nucleocapsid species. In addition to transcriptase and replicase functions, the viral L protein is also thought to have an endonuclease activity that cleaves cellular messenger RNAs (mRNAs) for the production of capped primers used to initiate transcription of viral mRNAs. As a result of this cap snatching, the mRNAs of hantaviruses are capped and contain nontemplated 5'-terminal extensions.[10]
The G1 (or Gn) and G2 (Gc) glycoproteins form hetero-oligomers and are then transported from the endoplasmic reticulum to the Golgi complex, where glycosylation is completed. The L protein produces nascent genomes by replication via a positive-sense RNA intermediate. Hantavirus virions are believed to assemble by association of nucleocapsids with glycoproteins embedded in the membranes of the Golgi, followed by budding into the Golgi cisternae. Nascent virions are then transported in secretory vesicles to the plasma membrane and released by exocytosis.[citation needed]
Pathogenesis
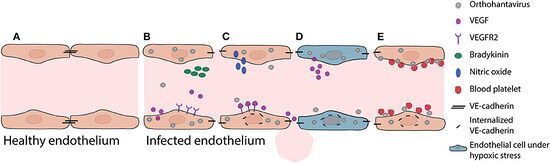
The pathogenesis of hantavirus infections is unclear as there is a lack of animal models to describe it (rats and mice do not seem to acquire severe disease). While the primary site of viral replication in the body is not known, in HFRS the main effect is in the blood vessels while in HPS most symptoms are associated with the lungs. In HFRS, there are increased vascular permeability and decreased blood pressure due to endothelial dysfunction and the most dramatic damage is seen in the kidneys, whereas in HPS, the lungs, spleen, and gall bladder are most affected. Early symptoms of HPS tend to present similarly to the flu (muscle aches, fever and fatigue) and usually appear around 2 to 3 weeks after exposure. Later stages of the disease (about 4 to 10 days after symptoms start) include difficulty breathing, shortness of breath and coughing.[12]
Evolution
Findings of significant congruence between phylogenies of hantaviruses and phylogenies of their rodent reservoirs have led to the theory that rodents, although infected by the virus, are not harmed by it because of long-standing hantavirus–rodent host coevolution,[13][14] although findings in 2008 led to new hypotheses regarding hantavirus evolution.[15][16]
Various hantaviruses have been found to infect multiple rodent species, and cases of cross-species transmission (host switching) have been recorded.[17][18][19] Additionally, rates of substitution based on nucleotide sequence data reveal that hantavirus clades and rodent subfamilies may not have diverged at the same time.[16][20] Furthermore, as of 2007 hantaviruses have been found in multiple species of non-rodent shrews and moles.[16][21][22][23]
Taking into account the inconsistencies in the theory of coevolution, it was proposed in 2009 that the patterns seen in hantaviruses in relation to their reservoirs could be attributed to preferential host switching directed by geographical proximity and adaptation to specific host types.[16] Another proposal from 2010 is that geographical clustering of hantavirus sequences may have been caused by an isolation-by-distance mechanism.[19] Upon comparison of the hantaviruses found in hosts of orders Rodentia and Eulipotyphla, it was proposed in 2011 that the hantavirus evolutionary history is a mix of both host switching and codivergence and that ancestral shrews or moles, rather than rodents, may have been the early original hosts of ancient hantaviruses.[21]
A Bayesian analysis in 2014 suggested a common origin for these viruses ~2000 years ago. The association with particular rodent families appears to have been more recent. The viruses carried by the subfamilies Arvicolinae and Murinae originated in Asia 500–700 years ago. These subsequently spread to Africa, Europe, North America and Siberia possibly carried by their hosts. The species infecting the subfamily Neotominae evolved 500–600 years ago in Central America and then spread toward North America. The species infecting Sigmodontinae evolved in Brazil 400 years ago. Their ancestors may have been a Neotominae-associated virus from northern South America.[24]
The evolution of shrew-borne hantaviruses appears to have involved natural occurrences of homologous recombination events and the reassortment of genome segments.[25] The evolution of Tula orthohantavirus carried by the European common vole also appears to have involved homologous recombination events.[26]
Taxonomy
Orthohantaviruses belong to the family Hantaviridae and members of both the genus and the family are called hantaviruses. The genus also belongs to the subfamily Mammantavirinae, the mammalian hantaviruses, with three other genera. Orthohantaviruses specifically are mammalian hantaviruses that are transmitted among rodents.[27] The genus has 36 recognized species as of 2019. The type species of the genus is the Hantaan orthohantavirus:[28]
- Andes orthohantavirus
- Asama orthohantavirus
- Asikkala orthohantavirus
- Bayou orthohantavirus
- Black Creek Canal orthohantavirus
- Bowe orthohantavirus
- Bruges orthohantavirus
- Cano Delgadito orthohantavirus
- Cao Bang orthohantavirus
- Choclo orthohantavirus
- Dabieshan orthohantavirus
- Dobrava-Belgrade orthohantavirus
- El Moro Canyon orthohantavirus
- Fugong orthohantavirus
- Fusong orthohantavirus
- Hantaan orthohantavirus
- Jeju orthohantavirus
- Kenkeme orthohantavirus
- Khabarovsk orthohantavirus
- Laguna Negra orthohantavirus
- Luxi orthohantavirus
- Maporal orthohantavirus
- Montano orthohantavirus
- Necocli orthohantavirus
- Oxbow orthohantavirus
- Prospect Hill orthohantavirus
- Puumala orthohantavirus
- Rockport orthohantavirus
- Sangassou orthohantavirus
- Seewis orhtohantavirus
- Seoul orthohantavirus
- Sin Nombre orthohantavirus
- Thailand orthohantavirus
- Tigray orthohantavirus
- Tula orthohantavirus
- Yakeshi orthohantavirus
See also
References
- ↑ "Virus Taxonomy: 2018b Release". International Committee on Taxonomy of Viruses (ICTV). March 2019. Archived from the original on 20 March 2020. Retrieved 18 March 2019.
- ↑ "ICTV Taxonomy all history: Orthohantavirus". International Committee on Taxonomy of Viruses (ICTV). Archived from the original on 28 January 2019. Retrieved 28 January 2019.
- ↑ 3.0 3.1 "Rodent-borne diseases". European Centre for Disease Prevention and Control. Archived from the original on 2018-06-23. Retrieved 2018-06-04.
- ↑ 4.0 4.1 Drebot, Jones S.; Grolla, A.; Safronetz, D.; Strong, J. E.; Kobinger, G.; Lindsay, R. L. (4 June 2015). Hantavirus pulmonary syndrome in Canada: An overview of clinical features, diagnostics, epidemiology and prevention. Canada Communicable Disease Report (Report). Vector-borne diseases in Canada. Vol. 41–06. Winnipeg, MB: National Microbiology Laboratory, Public Health Agency of Canada. p. 40. ISSN 1481-8531. Archived from the original on 19 July 2017. Retrieved 9 June 2015.
- ↑ Martinez VP, Bellomo C, San Juan J, Pinna D, Forlenza R, Elder M, Padula PJ (2005). "Person-to-person transmission of Andes virus". Emerging Infectious Diseases. 11 (12): 1848–1853. doi:10.3201/eid1112.050501. PMC 3367635. PMID 16485469.
- ↑ Gravinatti ML, Barbosa CM, Soares RM, Gregori F (2020). "Synanthropic rodents as virus reservoirs and transmitters". Rev Soc Bras Med Trop. 53: e20190486. doi:10.1590/0037-8682-0486-2019. PMC 7083353. PMID 32049206.
- ↑ 7.0 7.1 Klempa B (2018). "Reassortment events in the evolution of hantaviruses". Virus Genes. 54 (5): 638–646. doi:10.1007/s11262-018-1590-z. PMC 6153690. PMID 30047031.
- ↑ Elliott RM (1990). "Molecular biology of the Bunyaviridae". The Journal of General Virology. 71 (3): 501–522. doi:10.1099/0022-1317-71-3-501. PMID 2179464.
- ↑ Mir MA, Panganiban AT (2005). "The Hantavirus Nucleocapsid Protein Recognizes Specific Features of the Viral RNA Panhandle and is Altered in Conformation upon RNA Binding". Journal of Virology. 79 (3): 1824–1835. doi:10.1128/JVI.79.3.1824-1835.2005. PMC 544099. PMID 15650206.
- ↑ Garcin, D.; Lezzi, M.; Dobbs, M.; Elliott, R. M.; Schmaljohn, C.; Kang, C. Y.; Kolakofsky, D. (September 1995). "The 5' ends of Hantaan virus (Bunyaviridae) RNAs suggest a prime-and-realign mechanism for the initiation of RNA synthesis". Journal of Virology. 69 (9): 5754–5762. doi:10.1128/JVI.69.9.5754-5762.1995. ISSN 0022-538X. PMC 189436. PMID 7637020.
- ↑ Noack, Danny; Goeijenbier, Marco; Reusken, Chantal B. E. M.; Koopmans, Marion P. G.; Rockx, Barry H. G. (2020). "Orthohantavirus Pathogenesis and Cell Tropism". Frontiers in Cellular and Infection Microbiology. 10: 399. doi:10.3389/fcimb.2020.00399. ISSN 2235-2988.
- ↑ "Hantavirus: Canadian Lung Association". Canadian Lung Association. 26 November 2015. Archived from the original on 2 March 2011. Retrieved 23 April 2018.
- ↑ Plyusnin A, Vapalahti O, Vaheri A (1996). "Hantaviruses: genome structure, expression and evolution". J. Gen. Virol. 77 (11): 2677–2687. doi:10.1099/0022-1317-77-11-2677. PMID 8922460.
- ↑ Jackson AP, Charleston MA (2003). "A Cophylogenetic Perspective of RNA-Virus Evolution". Molecular Biology and Evolution. 21 (1): 45–57. doi:10.1093/molbev/msg232. PMID 12949128.
- ↑ Jonsson CB, Figueiredo LT, Vapalahti O (2010). "A Global Perspective on Hantavirus Ecology, Epidemiology, and Disease". Clinical Microbiology Reviews. 23 (2): 412–441. doi:10.1128/CMR.00062-09. PMC 2863364. PMID 20375360.
- ↑ 16.0 16.1 16.2 16.3 Ramsden C, Holmes EC, Charleston MA (2008). "Hantavirus Evolution in Relation to Its Rodent and Insectivore Hosts: No Evidence for Codivergence". Molecular Biology and Evolution. 26 (1): 143–153. doi:10.1093/molbev/msn234. PMID 18922760.
- ↑ Delfraro A, Tomé L, D'Elía G, Clara M, Achával F, Russi JC, Arbiza Rodonz JR (2008). "Juquitiba-like Hantavirus from 2 Nonrelated Rodent Species, Uruguay". Emerging Infectious Diseases. 14 (9): 1447–1451. doi:10.3201/eid1409.080455. PMC 2603116. PMID 18760017.
- ↑ Plyusnina A, Ibrahim IN, Plyusnin A (2009). "A newly recognized hantavirus in the Asian house rat (Rattus tanezumi) in Indonesia". Journal of General Virology. 90 (Pt 1): 205–209. doi:10.1099/Vir.0.006155-0. PMID 19088290.
- ↑ 19.0 19.1 Schmidt-Chanasit J, Essbauer S, Petraityte R, Yoshimatsu K, Tackmann K, Conraths FJ, Sasnauskas K, Arikawa J, Thomas A, Pfeffer M, Scharninghausen JJ, Splettstoesser W, Wenk M, Heckel G, Ulrich RG (2009). "Extensive Host Sharing of Central European Tula Virus". Journal of Virology. 84 (1): 459–474. doi:10.1128/Jvi.01226-09. PMC 2798396. PMID 19889769.
- ↑ Ramsden C, Melo FL, Figueiredo LM, Holmes EC, Zanotto PM (2008). "High Rates of Molecular Evolution in Hantaviruses". Molecular Biology and Evolution. 25 (7): 1488–1492. doi:10.1093/molbev/msn093. PMID 18417484.
- ↑ 21.0 21.1 Kang HJ, Bennett SN, Hope AG, Cook JA, Yanagihara R (2011). "Shared Ancestry between a Newfound Mole-Borne Hantavirus and Hantaviruses Harbored by Cricetid Rodents". Journal of Virology. 85 (15): 7496–7503. doi:10.1128/JVI.02450-10. PMC 3147906. PMID 21632770.
- ↑ Song JW, Baek LJ, Schmaljohn CS, Yanagihara R (2007). "Thottapalayam Virus, a Prototype Shrewborne Hantavirus". Emerging Infectious Diseases. 13 (7): 980–985. doi:10.3201/eid1307.070031. PMC 2254531. PMID 18214168.
- ↑ Song JW, Kang HJ, Song KJ, Truong TT, Bennett SN, Arai S, Truong NU, Yanagihara R (2007). "Newfound Hantavirus in Chinese Mole Shrew, Vietnam". Emerging Infectious Diseases. 13 (11): 1784–1787. doi:10.3201/eid1311.070492. PMC 2262106. PMID 18217572.
- ↑ Souza WM, Bello G, Amarilla AA, Alfonso HL, Aquino VH, Figueiredo LT (2014). "Phylogeography and evolutionary history of rodent-borne hantaviruses". Infect. Genet. Evol. 21: 198–204. doi:10.1016/j.meegid.2013.11.015. PMID 24287104.
- ↑ Lee SH, Kim WK, No JS, Kim JA, Kim JI, Gu SH, Kim HC, Klein TA, Park MS, Song JW (2017). "Dynamic Circulation and Genetic Exchange of a Shrew-borne Hantavirus, Imjin virus, in the Republic of Korea". Sci. Rep. 7: 44369. Bibcode:2017NatSR...744369L. doi:10.1038/srep44369. PMC 5353647. PMID 28295052.
- ↑ Sibold C, Meisel H, Krüger DH, Labuda M, Lysy J, Kozuch O, Pejcoch M, Vaheri A, Plyusnin A (1999). "Recombination in Tula hantavirus evolution: analysis of genetic lineages from Slovakia". J Virol. 73 (1): 667–75. doi:10.1128/jvi.73.1.667-675.1999. PMC 103873. PMID 9847372.
- ↑ "ICTV Taxonomy history: Mammantavirinae". talk.ictvonline.org. International Committee on Taxonomy of Viruses. Archived from the original on 28 August 2021. Retrieved 24 April 2020.
- ↑ "Virus Taxonomy: 2019 Release". talk.ictvonline.org. International Committee on Taxonomy of Viruses. Archived from the original on 20 March 2020. Retrieved 24 April 2020.
External links
- CDC's Hantavirus Fact Sheet (PDF) Archived 2017-08-29 at the Wayback Machine
- CDC's Hantavirus Technical Information Index page Archived 2012-01-18 at the Wayback Machine
- Virus Pathogen Database and Analysis Resource (ViPR): Hantaviridae Archived 2019-09-12 at the Wayback Machine
- Occurrences and deaths in North and South America
| Classification | |
|---|---|
| External resources |
- CS1: long volume value
- Articles with 'species' microformats
- All articles with unsourced statements
- Articles with unsourced statements from April 2018
- Articles with invalid date parameter in template
- Webarchive template wayback links
- Hantaviridae
- Hemorrhagic fevers
- Rodent-carried diseases
- Viral diseases
- Virus genera