Molecular motor
Molecular motors are natural (biological) or artificial molecular machines that are the essential agents of movement in living organisms. In general terms, a motor is a device that consumes energy in one form and converts it into motion or mechanical work; for example, many protein-based molecular motors harness the chemical free energy released by the hydrolysis of ATP in order to perform mechanical work.[1] In terms of energetic efficiency, this type of motor can be superior to currently available man-made motors. One important difference between molecular motors and macroscopic motors is that molecular motors operate in the thermal bath, an environment in which the fluctuations due to thermal noise are significant.
Examples
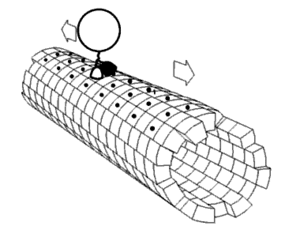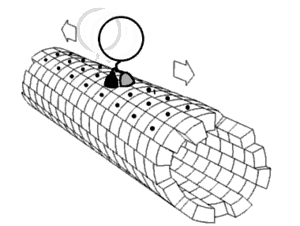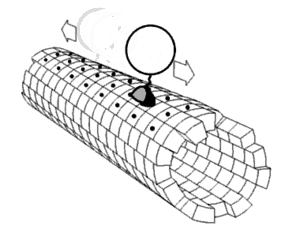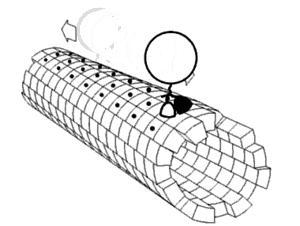
Some examples of biologically important molecular motors:[2]
- Cytoskeletal motors
- Myosins are responsible for muscle contraction, intracellular cargo transport, and producing cellular tension.
- Kinesin moves cargo inside cells away from the nucleus along microtubules, in anterograde transport.
- Dynein produces the axonemal beating of cilia and flagella and also transports cargo along microtubules towards the cell nucleus, in retrograde transport.
- Polymerisation motors
- Rotary motors:
- FoF1-ATP synthase family of proteins convert the chemical energy in ATP to the electrochemical potential energy of a proton gradient across a membrane or the other way around. The catalysis of the chemical reaction and the movement of protons are coupled to each other via the mechanical rotation of parts of the complex. This is involved in ATP synthesis in the mitochondria and chloroplasts as well as in pumping of protons across the vacuolar membrane.[3]
- The bacterial flagellum responsible for the swimming and tumbling of E. coli and other bacteria acts as a rigid propeller that is powered by a rotary motor. This motor is driven by the flow of protons across a membrane, possibly using a similar mechanism to that found in the Fo motor in ATP synthase.

- Nucleic acid motors:
- RNA polymerase transcribes RNA from a DNA template.[5]
- DNA polymerase turns single-stranded DNA into double-stranded DNA.[6]
- Helicases separate double strands of nucleic acids prior to transcription or replication. ATP is used.
- Topoisomerases reduce supercoiling of DNA in the cell. ATP is used.
- RSC and SWI/SNF complexes remodel chromatin in eukaryotic cells. ATP is used.
- SMC proteins responsible for chromosome condensation in eukaryotic cells.[7]
- Viral DNA packaging motors inject viral genomic DNA into capsids as part of their replication cycle, packing it very tightly.[8] Several models have been put forward to explain how the protein generates the force required to drive the DNA into the capsid. An alternative proposal is that, in contrast with all other biological motors, the force is not generated directly by the protein, but by the DNA itself.[9] In this model, ATP hydrolysis is used to drive protein conformational changes that alternatively dehydrate and rehydrate the DNA, cyclically driving it from B-DNA to A-DNA and back again. A-DNA is 23% shorter than B-DNA, and the DNA shrink/expand cycle is coupled to a protein-DNA grip/release cycle to generate the forward motion that propels DNA into the capsid.
- Enzymatic motors: The enzymes below have been shown to diffuse faster in the presence of their catalytic substrates, known as enhanced diffusion. They also have been shown to move directionally in a gradient of their substrates, known as chemotaxis. Their mechanisms of diffusion and chemotaxis are still debated. Possible mechanisms include solutal buoyancy, phoresis or conformational changes leading to change in effective diffusivity[10][11][12] and kinetic asymmetry.[13]
- Catalase
- Urease
- Aldolase
- Hexokinase
- Phosphoglucose isomerase
- Phosphofructokinase
- Glucose Oxidase
A recent study has also shown that certain enzymes, such as Hexokinase and Glucose Oxidase, are aggregating or fragmenting during catalysis. This changes their hydrodynamic size that can affect enhanced diffusion measurements.[14]
- Synthetic molecular motors have been created by chemists that yield rotation, possibly generating torque.[15]
Organelle and vesicle transport
There are two major families of molecular motors that transport organelles throughout the cell. These families include the dynein family and the kinesin family. Both have very different structures from one another and different ways of achieving a similar goal of moving organelles around the cell. These distances, though only few micrometers, are all preplanned out using microtubules.[16]
- Kinesin – These molecular motors always move towards the positive end of the cell
- Uses ATP hydrolysis during the process converting ATP to ADP
- This process consists of ...
- The "foot" of the motor binds using ATP, the "foot" proceeds a step, and then ADP comes off. This repeats itself until the destination has been reached
- This process consists of ...
- The kinesin family consists of a multitude of different motor types
- Kinesin-1 (Conventional)
- Kinesin-2 (Heterotrimeric)
- Kinesin-5 (Bipolar)
- Kinesin-13
- Uses ATP hydrolysis during the process converting ATP to ADP
- Dynein – These molecular motors always move towards the negative end of the cell
- Uses ATP hydrolysis during the process converting ATP to ADP
- Unlike kinesin, the dynein is structured in a different way which requires it to have different movement methods.
- One of these methods includes the power stroke, which allows the motor protein to "crawl" along the microtubule to its location.
- The structure of dynein consists of
- A Stem Containing
- A region that binds to dynactin
- Intermediate/light chains that will attach to the dynactin bonding region
- A Head
- A Stalk
- With a domain that will bind to the microtubule
- A Stem Containing
- These molecular motors tend to take the path of the microtubules. This is most likely due to the facts that the microtubules spring forth out of the centrosome and surround the entire volume of the cell. This in turn creates a "Rail system" of the whole cell and paths leading to its organelles.
Theoretical considerations
| Part of a series on |
| Microbial and microbot movement |
|---|
 |
| Microswimmers |
| Molecular motors |
Because the motor events are stochastic, molecular motors are often modeled with the Fokker–Planck equation or with Monte Carlo methods. These theoretical models are especially useful when treating the molecular motor as a Brownian motor.
Experimental observation
In experimental biophysics, the activity of molecular motors is observed with many different experimental approaches, among them:
- Fluorescent methods: fluorescence resonance energy transfer (FRET), fluorescence correlation spectroscopy (FCS), total internal reflection fluorescence (TIRF).
- Magnetic tweezers can also be useful for analysis of motors that operate on long pieces of DNA.
- Neutron spin echo spectroscopy can be used to observe motion on nanosecond timescales.
- Optical tweezers (not to be confused with molecular tweezers in context) are well-suited for studying molecular motors because of their low spring constants.
- Scattering techniques: single particle tracking based on dark field microscopy or interferometric scattering microscopy (iSCAT)
- Single-molecule electrophysiology can be used to measure the dynamics of individual ion channels.
Many more techniques are also used. As new technologies and methods are developed, it is expected that knowledge of naturally occurring molecular motors will be helpful in constructing synthetic nanoscale motors.
Non-biological
Recently, chemists and those involved in nanotechnology have begun to explore the possibility of creating molecular motors de novo. [17] These synthetic molecular motors currently suffer many limitations that confine their use to the research laboratory. However, many of these limitations may be overcome as our understanding of chemistry and physics at the nanoscale increases. One step toward understanding nanoscale dynamics was made with the study of catalyst diffusion in the Grubb's catalyst system.[18] Other systems like the nanocars, while not technically motors, are also illustrative of recent efforts towards synthetic nanoscale motors.
Other non-reacting molecules can also behave as motors. This has been demonstrated by using dye molecules that move directionally in gradients of polymer solution through favorable hydrophobic interactions.[19] Another recent study has shown that dye molecules, hard and soft colloidal particles are able to move through gradient of polymer solution through excluded volume effects.[20]
See also
- Brownian motor
- Brownian ratchet
- Cytoskeleton
- Molecular machines
- Molecular mechanics
- Molecular propeller
- Motor proteins
- Nanomotor
- Protein dynamics
- Synthetic molecular motors
References
- ^ Bustamante C, Chemla YR, Forde NR, Izhaky D (2004). "Mechanical processes in biochemistry". Annual Review of Biochemistry. 73: 705–48. doi:10.1146/annurev.biochem.72.121801.161542. PMID 15189157. S2CID 28061339.
- ^ Nelson P, Radosavljevic M, Bromberg S (2004). Biological physics. Freeman.
- ^ Tsunoda SP, Aggeler R, Yoshida M, Capaldi RA (January 2001). "Rotation of the c subunit oligomer in fully functional F1Fo ATP synthase". Proceedings of the National Academy of Sciences of the United States of America. 98 (3): 898–902. Bibcode:2001PNAS...98..898T. doi:10.1073/pnas.031564198. PMC 14681. PMID 11158567.
- ^ Palma CA, Björk J, Rao F, Kühne D, Klappenberger F, Barth JV (August 2014). "Topological dynamics in supramolecular rotors". Nano Letters. 14 (8): 4461–8. Bibcode:2014NanoL..14.4461P. doi:10.1021/nl5014162. PMID 25078022.
- ^ Dworkin J, Losick R (October 2002). "Does RNA polymerase help drive chromosome segregation in bacteria?". Proceedings of the National Academy of Sciences of the United States of America. 99 (22): 14089–94. Bibcode:2002PNAS...9914089D. doi:10.1073/pnas.182539899. PMC 137841. PMID 12384568.
- ^ Hubscher U, Maga G, Spadari S (2002). "Eukaryotic DNA polymerases". Annual Review of Biochemistry. 71: 133–63. doi:10.1146/annurev.biochem.71.090501.150041. PMID 12045093. S2CID 26171993.
- ^ Peterson CL (November 1994). "The SMC family: novel motor proteins for chromosome condensation?". Cell. 79 (3): 389–92. doi:10.1016/0092-8674(94)90247-X. PMID 7954805. S2CID 28364947.
- ^ Smith DE, Tans SJ, Smith SB, Grimes S, Anderson DL, Bustamante C (October 2001). "The bacteriophage straight phi29 portal motor can package DNA against a large internal force". Nature. 413 (6857): 748–52. Bibcode:2001Natur.413..748S. doi:10.1038/35099581. PMID 11607035. S2CID 4424168.
- ^ Harvey SC (January 2015). "The scrunchworm hypothesis: transitions between A-DNA and B-DNA provide the driving force for genome packaging in double-stranded DNA bacteriophages". Journal of Structural Biology. 189 (1): 1–8. doi:10.1016/j.jsb.2014.11.012. PMC 4357361. PMID 25486612.
- ^ Zhao X, Gentile K, Mohajerani F, Sen A (October 2018). "Powering Motion with Enzymes". Accounts of Chemical Research. 51 (10): 2373–2381. doi:10.1021/acs.accounts.8b00286. PMID 30256612. S2CID 52845451.
- ^ Ghosh S, Somasundar A, Sen A (2021-03-10). "Enzymes as Active Matter". Annual Review of Condensed Matter Physics. 12 (1): 177–200. Bibcode:2021ARCMP..12..177G. doi:10.1146/annurev-conmatphys-061020-053036. S2CID 229411011.
- ^ Zhang Y, Hess H (June 2019). "Enhanced Diffusion of Catalytically Active Enzymes". ACS Central Science. 5 (6): 939–948. doi:10.1021/acscentsci.9b00228. PMC 6598160. PMID 31263753.
- ^ Mandal, Niladri Sekhar; Sen, Ayusman; Astumian, R. Dean (2023-03-15). "Kinetic Asymmetry versus Dissipation in the Evolution of Chemical Systems as Exemplified by Single Enzyme Chemotaxis". Journal of the American Chemical Society. 145 (10): 5730–5738. arXiv:2206.05626. doi:10.1021/jacs.2c11945. ISSN 0002-7863. PMID 36867055. S2CID 249625518.
- ^ Gentile, Kayla; Bhide, Ashlesha; Kauffman, Joshua; Ghosh, Subhadip; Maiti, Subhabrata; Adair, James; Lee, Tae-Hee; Sen, Ayusman (2021-09-22). "Enzyme aggregation and fragmentation induced by catalysis relevant species". Physical Chemistry Chemical Physics. 23 (36): 20709–20717. Bibcode:2021PCCP...2320709G. doi:10.1039/D1CP02966E. ISSN 1463-9084. PMID 34516596. S2CID 237507756.
- ^ Kay, Euan R.; Leigh, David A.; Zerbetto, Francesco (January 2007). "Synthetic Molecular Motors and Mechanical Machines". Angewandte Chemie International Edition. 46 (1–2): 72–191. doi:10.1002/anie.200504313. PMID 17133632.
- ^ Lodish H, Berk A, Kaiser CA, Krieger M, Bretscher A, Ploegh H, Amon A, Martin KC (2014). Molecular Cell Biology (8th ed.). New York, NY: w.h.freeman, Macmillan Learning. ISBN 978-1-4641-8339-3.
- ^ Korosec, Chapin S.; Unksov, Ivan N.; Surendiran, Pradheebha; Lyttleton, Roman; Curmi, Paul M. G.; Angstmann, Christopher N.; Eichhorn, Ralf; Linke, Heiner; Forde, Nancy R. (2024-02-23). "Motility of an autonomous protein-based artificial motor that operates via a burnt-bridge principle". Nature Communications. 15 (1511): 1511. Bibcode:2024NatCo..15.1511K. doi:10.1038/s41467-024-45570-y. PMC 10891099. PMID 38396042.
- ^ Dey KK, Pong FY, Breffke J, Pavlick R, Hatzakis E, Pacheco C, Sen A (January 2016). "Dynamic Coupling at the Ångström Scale". Angewandte Chemie. 55 (3): 1113–7. Bibcode:2016AngCh.128.1125D. doi:10.1002/ange.201509237. PMID 26636667.
- ^ Guha R, Mohajerani F, Collins M, Ghosh S, Sen A, Velegol D (November 2017). "Chemotaxis of Molecular Dyes in Polymer Gradients in Solution". Journal of the American Chemical Society. 139 (44): 15588–15591. doi:10.1021/jacs.7b08783. PMID 29064685.
- ^ Collins M, Mohajerani F, Ghosh S, Guha R, Lee TH, Butler PJ, et al. (August 2019). "Nonuniform Crowding Enhances Transport". ACS Nano. 13 (8): 8946–8956. doi:10.1021/acsnano.9b02811. PMID 31291087. S2CID 195879481.
External links
- MBInfo - Molecular Motor Activity
- MBInfo - Cytoskeleton-dependent MBInfo - Intracellular Transport
- Cymobase - A database for cytoskeletal and motor protein sequence information
- Jonathan Howard (2001), Mechanics of motor proteins and the cytoskeleton. ISBN 9780878933334