CRISPR gene editing

CRISPR gene editing (pronounced /ˈkrɪspər/ "crisper") is a genetic engineering technique in molecular biology by which the genomes of living organisms may be modified. It is based on a simplified version of the bacterial CRISPR-Cas9 antiviral defense system. By delivering the Cas9 nuclease complexed with a synthetic guide RNA (gRNA) into a cell, the cell's genome can be cut at a desired location, allowing existing genes to be removed and/or new ones added in vivo.[1]
The technique is considered highly significant in biotechnology and medicine as it enables editing genomes in vivo very precisely, cheaply, and easily. It can be used in the creation of new medicines, agricultural products, and genetically modified organisms, or as a means of controlling pathogens and pests. It also has possibilities in the treatment of inherited genetic diseases as well as diseases arising from somatic mutations such as cancer. However, its use in human germline genetic modification is highly controversial. The development of the technique earned Jennifer Doudna and Emmanuelle Charpentier the Nobel Prize in Chemistry in 2020.[2][3] The third researcher group that shared the Kavli Prize for the same discovery,[4] led by Virginijus Šikšnys, was not awarded the Nobel prize.[5][6][7]
Working like genetic scissors, the Cas9 nuclease opens both strands of the targeted sequence of DNA to introduce the modification by one of two methods. Knock-in mutations, facilitated via homology directed repair (HDR), is the traditional pathway of targeted genomic editing approaches.[1] This allows for the introduction of targeted DNA damage and repair. HDR employs the use of similar DNA sequences to drive the repair of the break via the incorporation of exogenous DNA to function as the repair template.[1] This method relies on the periodic and isolated occurrence of DNA damage at the target site in order for the repair to commence. Knock-out mutations caused by CRISPR-Cas9 result from the repair of the double-stranded break by means of non-homologous end joining (NHEJ) or POLQ/ polymerase theta-mediated end-joining (TMEJ). These end-joining pathways can often result in random deletions or insertions at the repair site, which may disrupt or alter gene functionality. Therefore, genomic engineering by CRISPR-Cas9 gives researchers the ability to generate targeted random gene disruption. Because of this, the precision of genome editing is a great concern. Genomic editing leads to irreversible changes to the genome.
While genome editing in eukaryotic cells has been possible using various methods since the 1980s, the methods employed had proved to be inefficient and impractical to implement on a large scale. With the discovery of CRISPR and specifically the Cas9 nuclease molecule, efficient and highly selective editing becomes possible. Cas9 derived from the bacterial species Streptococcus pyogenes has facilitated targeted genomic modification in eukaryotic cells by allowing for a reliable method of creating a targeted break at a specific location as designated by the crRNA and tracrRNA guide strands.[8] The ease with which researchers can insert Cas9 and template RNA in order to silence or cause point mutations at specific loci has proved invaluable to the quick and efficient mapping of genomic models and biological processes associated with various genes in a variety of eukaryotes. Newly engineered variants of the Cas9 nuclease have been developed that significantly reduce off-target activity.[9]
CRISPR-Cas9 genome editing techniques have many potential applications, including in medicine and agriculture. The use of the CRISPR-Cas9-gRNA complex for genome editing[10] was the AAAS's choice for Breakthrough of the Year in 2015.[11] Many bioethical concerns have been raised about the prospect of using CRISPR for germline editing, especially in human embryos.[12]
In 2023, the first drug making use of CRISPR gene editing, Casgevy, was approved for use in the United Kingdom to cure sickle-cell disease and beta thalassemia.[13][14]
History
Other methods
In the early 2000s, German researchers began developing zinc finger nucleases (ZFNs), synthetic proteins whose DNA-binding domains enable them to create double-stranded breaks in DNA at specific points. ZFNs has a higher precision and the advantage of being smaller than Cas9, but ZFNs are not as commonly used as CRISPR-based methods. In 2010, synthetic nucleases called transcription activator-like effector nucleases (TALENs) provided an easier way to target a double-stranded break to a specific location on the DNA strand. Both zinc finger nucleases and TALENs require the design and creation of a custom protein for each targeted DNA sequence, which is a much more difficult and time-consuming process than that of designing guide RNAs. CRISPRs are much easier to design because the process requires synthesizing only a short RNA sequence, a procedure that is already widely used for many other molecular biology techniques (e.g. creating oligonucleotide primers).[15]
Whereas methods such as RNA interference (RNAi) do not fully suppress gene function, CRISPR, ZFNs, and TALENs provide full irreversible gene knockout.[16] CRISPR can also target several DNA sites simultaneously simply by introducing different gRNAs. In addition, the costs of employing CRISPR are relatively low.[16][17][18]
Discovery
In 2012 Jennifer Doudna, Emmanuelle Charpentier, Martin Jínek, Krzysztof Chyliński, Ines Fonfara and Michael Hauer published their finding that CRISPR-Cas9 could be programmed with RNA to edit genomic DNA, now considered one of the most significant discoveries in the history of biology.[19][20]
Patents and commercialization
As of November 2013[update], SAGE Labs (part of Horizon Discovery group) had exclusive rights from one of those companies to produce and sell genetically engineered rats and non-exclusive rights for mouse and rabbit models.[21] By 2015[update], Thermo Fisher Scientific had licensed intellectual property from ToolGen to develop CRISPR reagent kits.[22]
As of December 2014[update], patent rights to CRISPR were contested. Several companies formed to develop related drugs and research tools.[23] As companies ramped up financing, doubts as to whether CRISPR could be quickly monetized were raised.[24] In 2014, Feng Zhang of the Broad Institute of MIT and Harvard and nine others were awarded US patent number 8,697,359[25] over the use of CRISPR–Cas9 gene editing in eukaryotes. Although Charpentier and Doudna (referred to as CVC) were credited for the conception of CRISPR, the Broad Institute was the first to achieve a "reduction to practice" according to patent judges Sally Gardner Lane, James T. Moore and Deborah Katz.[26]
The first set of patents was awarded to the Broad team in 2015, prompting attorneys for the CVC group to request the first interference proceeding.[27] In February 2017 the US Patent Office ruled on a patent interference case brought by University of California with respect to patents issued to the Broad Institute, and found that the Broad patents, with claims covering the application of CRISPR-Cas9 in eukaryotic cells, were distinct from the inventions claimed by University of California.[28][29][30]
Shortly after, University of California filed an appeal of this ruling.[31][32] In 2019 the second interference dispute was opened. This was in response to patent applications made by CVC that required the appeals board to determine the original inventor of the technology. The USPTO ruled in March 2022 against UC, stating that the Broad Institute were first to file. The decision affected many of the licensing agreements for the CRISPR editing technology that was licensed from UC Berkeley. UC stated its intent to appeal the USPTO's ruling.[33]
Recent events
In March 2017, the European Patent Office (EPO) announced its intention to allow claims for editing all types of cells to Max-Planck Institute in Berlin, University of California, and University of Vienna,[34][35] and in August 2017, the EPO announced its intention to allow CRISPR claims in a patent application that MilliporeSigma had filed.[34] As of August 2017[update] the patent situation in Europe was complex, with MilliporeSigma, ToolGen, Vilnius University, and Harvard contending for claims, along with University of California and Broad.[36]
In July 2018, the ECJ ruled that gene editing for plants was a sub-category of GMO foods and therefore that the CRISPR technique would henceforth be regulated in the European Union by their rules and regulations for GMOs.[37]
In February 2020, a US trial showed safe CRISPR gene editing on three cancer patients.[38]
In October 2020, researchers Emmanuelle Charpentier and Jennifer Doudna were awarded the Nobel Prize in Chemistry for their work in this field.[39][40] They made history as the first two women to share this award without a male contributor.[41][5]
In June 2021, the first, small clinical trial of intravenous CRISPR gene editing in humans concludes with promising results.[42][43]
In September 2021, the first CRISPR-edited food has gone on public sale in Japan. Tomatoes were genetically modified for around five times the normal amount of possibly calming[44] GABA.[45] CRISPR was first applied in tomatoes in 2014.[46]
In December 2021, it was reported that the first CRISPR-gene-edited marine animal/seafood and second set of CRISPR-edited food has gone on public sale in Japan: two fish of which one species grows to twice the size of natural specimens due to disruption of leptin, which controls appetite, and the other grows to 1.2 the natural average size with the same amount of food due to disabled myostatin, which inhibits muscle growth.[47][48][49]
A 2022 study has found that knowing more about CRISPR tomatoes had a strong effect on the participants' preference. "Almost half of the 32 participants from Germany who are scientists demonstrated constant choices, while the majority showed increased willingness to buy CRISPR tomatoes, mostly non-scientists."[50][51]
The UC Berkeley announced in May 2021 their intent to auction non-fungible tokens of both the patent for CRISPR gene editing as well as cancer immunotherapy. The university would in this case retain ownership of the patents, however.[52][53] 85 % of funds gathered through the sale of the collection named The Fourth Pillar were to be used to finance research.[54][55] It sold in June 2022 for 22 Ether, which was around US$54,000 at the time.[56]
In a 2023 world first, the United Kingdom's Medicines and Healthcare products Regulatory Agency (MHRA) approved the use of the first drug based on CRISPR gene editing, Casgevy, to treat sickle-cell anemia and beta thalassemia. Casgevy, or exagamglogene autotemcel, directly acts on the genes of the stem cells inside the patient's bones, having them produce healthy red blood cells. This treatment thus avoids the need for regular, costly blood transfusions.[13][14]
Genome engineering
-
DNA repair after double-stranded break
-
Overview of CRISPR-Cas9 plasmid construction
CRISPR-Cas9 genome editing is carried out with a Type II CRISPR system. When utilized for genome editing, this system includes a ribonucleoprotein (RNP), consisting of Cas9, crRNA, and tracrRNA, along with an optional DNA repair template.
Major components
| Component | Function |
|---|---|
| crRNA | Contains the guide RNA that locates the correct segment of host DNA along with a region that binds to tracrRNA (generally in a hairpin loop form), forming an active complex. |
| tracrRNA | Binds to crRNA and forms an active complex. |
| sgRNA | Single-guide RNAs are a combined RNA consisting of a tracrRNA and at least one crRNA. |
| Cas9 (most commonly) | An enzyme whose active form is able to modify DNA. Many variants exist with different functions (i.e. single-strand nicking, double-strand breaking, DNA binding) due to each enzyme's DNA site recognition function. |
| Repair template | DNA molecule used as a template in the host cell's DNA repair process, allowing insertion of a specific DNA sequence into the host segment broken by Cas9. |

CRISPR-Cas9 often employs plasmids, that code for the RNP components, to transfect the target cells, or the RNP is assembled before addition to the cells via nucleofection.[57]
The main components of this plasmid are displayed in the image and listed in the table. The crRNA is uniquely designed for each application, as this is the sequence that Cas9 uses to identify and directly bind to specific sequences within the host cell's DNA. The crRNA must bind only where editing is desired. The repair template is also uniquely designed for each application, as it must complement to some degree the DNA sequences on either side of the cut and also contain whatever sequence is desired for insertion into the host genome.
Multiple crRNAs and the tracrRNA can be packaged together to form a single-guide RNA (sgRNA).[58]
This sgRNA can be included alongside the gene that codes for the Cas9 protein and made into a plasmid in order to be transfected into cells. Many online tools are available to aid in designing effective sgRNA sequences.[59][60]
Alternatives to Cas9

Alternative proteins to Cas9 include the following:
| Protein | Main use / characteristics | Year/s |
|---|---|---|
| Cas12 | Cas12a is smaller and simpler than Cas9; Cas12b i.a. for plant genome engineering[61][62] | |
| Cas13 | for RNA editing[63] | |
| Cas3[64][65] | Creates a single-stranded wide gap[66] | 2019 |
| CasMINI | About twice as compact as the more commonly used Cas9 and Cas12a.[67][68] | 2021 |
| SuperFi-Cas9 | More accurate without a slow down in speed[69][70] | 2022 |
| Cas7-11 | Better suited for therapeutic RNA editing than Cas13[71] | 2022 |
| Chromosome-templated DNA repair | Such a method is only applicable to organisms whose matching chromosome has the desired gene/s.
|
2022 |
Structure
CRISPR-Cas9 offers a high degree of fidelity and relatively simple construction. It depends on two factors for its specificity: the target sequence and the protospacer adjacent motif (PAM) sequence. The target sequence is 20 bases long as part of each CRISPR locus in the crRNA array.[57] A typical crRNA array has multiple unique target sequences. Cas9 proteins select the correct location on the host's genome by utilizing the sequence to bond with base pairs on the host DNA. The sequence is not part of the Cas9 protein and as a result is customizable and can be independently synthesized.[74][75]
The PAM sequence on the host genome is recognized by Cas9. Cas9 cannot be easily modified to recognize a different PAM sequence. However, this is ultimately not too limiting, as it is typically a very short and nonspecific sequence that occurs frequently at many places throughout the genome (e.g. the SpCas9 PAM sequence is 5'-NGG-3' and in the human genome occurs roughly every 8 to 12 base pairs).[57]
Once these sequences have been assembled into a plasmid and transfected into cells, the Cas9 protein with the help of the crRNA finds the correct sequence in the host cell's DNA and – depending on the Cas9 variant – creates a single- or double-stranded break at the appropriate location in the DNA.[76]
Properly spaced single-stranded breaks in the host DNA can trigger homology directed repair, which is less error-prone than the non-homologous end joining that typically follows a double-stranded break. Providing a DNA repair template allows for the insertion of a specific DNA sequence at an exact location within the genome. The repair template should extend 40 to 90 base pairs beyond the Cas9-induced DNA break.[57] The goal is for the cell's native HDR process to utilize the provided repair template and thereby incorporate the new sequence into the genome. Once incorporated, this new sequence is now part of the cell's genetic material and passes into its daughter cells. Combined transient inhibition of NHEJ and TMEJ by a small molecule and siRNAs can increase HDR efficiency to up to 93% and simultaneously prevent off-target editing.[77]
Delivery
Delivery of Cas9, sgRNA, and associated complexes into cells can occur via viral and non-viral systems. Electroporation of DNA, RNA, or ribonucleocomplexes is a common technique, though it can result in harmful effects on the target cells.[78] Chemical transfection techniques utilizing lipids and peptides have also been used to introduce sgRNAs in complex with Cas9 into cells.[79][80] Nanoparticle-based delivery has also been used for transfection.[81] Types of cells that are more difficult to transfect (e.g., stem cells, neurons, and hematopoietic cells) require more efficient delivery systems, such as those based on lentivirus (LVs), adenovirus (AdV), and adeno-associated virus (AAV).[82][83][84]
Efficiency of CRISPR-Cas9 has been found to greatly increase when various components of the system including the entire CRISPR/Cas9 structure to Cas9-gRNA complexes delivered in assembled form rather than using transgenics.[85][86] This has found particular value in genetically modified crops for mass commercialization.[87][88] Since the host's replication machinery is not needed to produce these proteins, the chance of the recognizing sequence of the sgRNA is almost none, decreasing the chance of off-target effects.[81]
Controlled genome editing
Further improvements and variants of the CRISPR-Cas9 system have focused on introducing more control into its use. Specifically, the research aimed at improving this system includes improving its specificity, its efficiency, and the granularity of its editing power. Techniques can further be divided and classified by the component of the system they modify. These include using a different variants or novel creations of the Cas protein, using an altogether different effector protein, modifying the sgRNA, or using an algorithmic approach to identify existing optimal solutions.
Specificity is an important aspect to improve the CRISPR-Cas9 system because the off-target effects it generates have serious consequences for the genome of the cell and invokes caution for its use. Minimizing off-target effects is thus maximizing the safety of the system. Novel variations of Cas9 proteins that increase specificity include effector proteins with comparable efficiency and specificity to the original SpCas9 that are able to target the previously untargetable sequences and a variant that has virtually no off-target mutations.[89][90] Research has also been conducted in engineering new Cas9 proteins, including some that partially replace RNA nucleotides in crRNA with DNA and a structure-guided Cas9 mutant generating procedure that all had reduced off-target effects.[91][92] Iteratively truncated sgRNAs and highly stabilized gRNAs have been shown to also decrease off-target effects.[93][94] Computational methods including machine learning have been used to predict the affinity of and create unique sequences for the system to maximize specificity for given targets.[95][96]
Several variants of CRISPR-Cas9 allow gene activation or genome editing with an external trigger such as light or small molecules.[97][98][99] These include photoactivatable CRISPR systems developed by fusing light-responsive protein partners with an activator domain and a dCas9 for gene activation,[100][101] or by fusing similar light-responsive domains with two constructs of split-Cas9,[102][103] or by incorporating caged unnatural amino acids into Cas9,[104] or by modifying the guide RNAs with photocleavable complements for genome editing.[105]
Methods to control genome editing with small molecules include an allosteric Cas9, with no detectable background editing, that will activate binding and cleavage upon the addition of 4-hydroxytamoxifen (4-HT),[97] 4-HT responsive intein-linked Cas9,[106] or a Cas9 that is 4-HT responsive when fused to four ERT2 domains.[107] Intein-inducible split-Cas9 allows dimerization of Cas9 fragments[108] and rapamycin-inducible split-Cas9 system developed by fusing two constructs of split-Cas9 with FRB and FKBP fragments.[109] Other studies have been able to induce transcription of Cas9 with a small molecule, doxycycline.[110][111] Small molecules can also be used to improve homology directed repair,[112] often by inhibiting the non-homologous end joining pathway and/or the theta-mediated end-joining pathway.[113][114] A system with the Cpf1 effector protein was created that is induced by small molecules VE-822 and AZD-7762.[115] These systems allow conditional control of CRISPR activity for improved precision, efficiency, and spatiotemporal control. Spatiotemporal control is a form of removing off-target effects—only certain cells or parts of the organism may need to be modified, and thus light or small molecules can be used as a way to conduct this. Efficiency of the CRISPR-Cas9 system is also greatly increased by proper delivery of the DNA instructions for creating the proteins and necessary reagents.[115]
CRISPR also utilizes single base-pair editing proteins to create specific edits at one or two bases in the target sequence. CRISPR/Cas9 was fused with specific enzymes that initially could only change C to T and G to A mutations and their reverse. This was accomplished eventually without requiring any DNA cleavage.[116][117][118] With the fusion of another enzyme, the base editing CRISPR-Cas9 system can also edit C to G and its reverse.[119]
CRISPR screening
The clustered regularly interspaced short palindrome repeats (CRISPR)/Cas9 system is a gene-editing technology that can induce double-strand breaks (DSBs) anywhere guide ribonucleic acids (gRNA) can bind with the protospacer adjacent motif (PAM) sequence.[120] Single-strand nicks can also be induced by Cas9 active-site mutants,[121] also known as Cas9 nickases.[122] By simply changing the sequence of gRNA, the Cas9-endonuclease can be delivered to a gene of interest and induce DSBs.[123] The efficiency of Cas9-endonuclease and the ease by which genes can be targeted led to the development of CRISPR-knockout (KO) libraries both for mouse and human cells, which can cover either specific gene sets of interest or the whole-genome.[124][125] CRISPR screening helps scientist to create a systematic and high-throughput genetic perturbation within live model organisms. This genetic perturbation is necessary for fully understanding gene function and epigenetic regulation.[126] The advantage of pooled CRISPR libraries is that more genes can be targeted at once.
Knock-out libraries are created in a way to achieve equal representation and performance across all expressed gRNAs and carry an antibiotic or fluorescent selection marker that can be used to recover transduced cells.[120] There are two plasmid systems in CRISPR/Cas9 libraries. First, is all in one plasmid, where sgRNA and Cas9 are produced simultaneously in a transfected cell. Second, is a two-vector system: sgRNA and Cas9 plasmids are delivered separately.[126] It is important to deliver thousands of unique sgRNAs-containing vectors to a single vessel of cells by viral transduction at low multiplicity of infection (MOI, typically at 0.1–0.6), it prevents the probability that an individual cell clone will get more than one type of sgRNA otherwise it can lead to incorrect assignment of genotype to phenotype.[124]
Once pooled library is prepared it is necessary to carry out a deep sequencing (NGS, next generation sequencing) of PCR-amplifed plasmid DNA in order to reveal abundance of sgRNAs. Cells of interest can be consequentially infected by the library and then selected according to the phenotype. There are 2 types of selection: negative and positive. By negative selection dead or slow growing cells are efficiently detected. It can identify survival-essential genes, which can be further serve as candidates for molecularly targeted drugs. On the other hand, positive selection gives a collection of growth-advantage acquired populations by random mutagenesis.[120] After selection genomic DNA is collected and sequenced by NGS. Depletion or enrichment of sgRNAs is detected and compared to the original sgRNA library, annotated with the target gene that sgRNA corresponds to. Statistical analysis then identify genes that are significantly likely to be relevant to the phenotype of interest.[124]
| Library | ID | Species | PI | Genes targeted | gRNAs per gene | Total gRNAs |
|---|---|---|---|---|---|---|
| Bassik Mouse CRISPR Knockout Library | 1000000121–1000000130 | Mouse | Bassik | Varies (~23,000 in total) | ~10 | Varies |
| Mouse Tumor Suppressor Gene CRISPR Knockout Library | 113584 EFS backbone
113585 TBG backbone |
Mouse | Chen | 56 | ~4 | 286 |
| Brie mouse genome-wide library | 73632 (1 plasmid)
73633 (2 plasmid) |
Mouse | Doench and Root | 19,674 | 4 | 78,637 |
| Bassik Human CRISPR Knockout Library | 101926–101934 | Human | Bassik | Varies (~20,500 in total) | ~10 | Varies |
| Brunello human genome-wide library | 73179 (1 plasmid)
73178 (2 plasmid) |
Human | Doench and Root | 19,114 | 4 | 76,441 |
| Mini-human AsCpf1-based Human Genome-wide Knockout Library | 130630 | Human | Draetta | 16,977 | 3–4 | 17,032 arrays |
Apart from knock-out there are also knock-down (CRISPRi) and activation (CRISPRa) libraries, which using the ability of proteolytically deactivated Cas9-fusion proteins (dCas9) to bind target DNA, which means that gene of interest is not cut but is over-expressed or repressed. It made CRISPR/Cas9 system even more interesting in gene editing. Inactive dCas9 protein modulate gene expression by targeting dCas9-repressors or activators toward promoter or transcriptional start sites of target genes. For repressing genes Cas9 can be fused to KRAB effector domain that makes complex with gRNA, whereas CRISPRa utilizes dCas9 fused to different transcriptional activation domains, which are further directed by gRNA to promoter regions to upregulate expression.[128][129][130]
Applications
Disease models
Cas9 genomic modification has allowed for the quick and efficient generation of transgenic models within the field of genetics. Cas9 can be easily introduced into the target cells along with sgRNA via plasmid transfection in order to model the spread of diseases and the cell's response to and defense against infection.[131] The ability of Cas9 to be introduced in vivo allows for the creation of more accurate models of gene function and mutation effects, all while avoiding the off-target mutations typically observed with older methods of genetic engineering.
The CRISPR and Cas9 revolution in genomic modeling does not extend only to mammals. Traditional genomic models such as Drosophila melanogaster, one of the first model organisms, have seen further refinement in their resolution with the use of Cas9.[131] Cas9 uses cell-specific promoters allowing a controlled use of the Cas9. Cas9 is an accurate method of treating diseases due to the targeting of the Cas9 enzyme only affecting certain cell types. The cells undergoing the Cas9 therapy can also be removed and reintroduced to provide amplified effects of the therapy.[132]
CRISPR-Cas9 can be used to edit the DNA of organisms in vivo and to eliminate individual genes or even entire chromosomes from an organism at any point in its development. Chromosomes that have been successfully deleted in vivo using CRISPR techniques include the Y chromosome and X chromosome of adult lab mice and human chromosomes 14 and 21, in embryonic stem cell lines and aneuploid mice respectively. This method might be useful for treating genetic disorders caused by abnormal numbers of chromosomes, such as Down syndrome and intersex disorders.[133]
Successful in vivo genome editing using CRISPR-Cas9 has been shown in numerous model organisms, including Escherichia coli,[134] Saccharomyces cerevisiae,[135] [136]Candida albicans, Methanosarcina acetivorans,[137][138] Caenorhabditis elegans,[139] Arabidopsis spp.,[140] Danio rerio,[141] and Mus musculus.[142][143] Successes have been achieved in the study of basic biology, in the creation of disease models,[139][144] and in the experimental treatment of disease models.[145]
Concerns have been raised that off-target effects (editing of genes besides the ones intended) may confound the results of a CRISPR gene editing experiment (i.e. the observed phenotypic change may not be due to modifying the target gene, but some other gene). Modifications to CRISPR have been made to minimize the possibility of off-target effects. Orthogonal CRISPR experiments are often recommended to confirm the results of a gene editing experiment.[146][147]
CRISPR simplifies the creation of genetically modified organisms for research which mimic disease or show what happens when a gene is knocked down or mutated. CRISPR may be used at the germline level to create organisms in which the targeted gene is changed everywhere (i.e. in all cells/tissues/organs of a multicellular organism), or it may be used in non-germline cells to create local changes that only affect certain cell populations within the organism.[148][149][150]
CRISPR can be utilized to create human cellular models of disease.[151] For instance, when applied to human pluripotent stem cells, CRISPR has been used to introduce targeted mutations in genes relevant to polycystic kidney disease (PKD) and focal segmental glomerulosclerosis (FSGS).[152] These CRISPR-modified pluripotent stem cells were subsequently grown into human kidney organoids that exhibited disease-specific phenotypes. Kidney organoids from stem cells with PKD mutations formed large, translucent cyst structures from kidney tubules. The cysts were capable of reaching macroscopic dimensions, up to one centimeter in diameter.[153] Kidney organoids with mutations in a gene linked to FSGS developed junctional defects between podocytes, the filtering cells affected in that disease. This was traced to the inability of podocytes to form microvilli between adjacent cells.[154] Importantly, these disease phenotypes were absent in control organoids of identical genetic background, but lacking the CRISPR modifications.[152]
A similar approach was taken to model long QT syndrome in cardiomyocytes derived from pluripotent stem cells.[155] These CRISPR-generated cellular models, with isogenic controls, provide a new way to study human disease and test drugs.
Biomedicine
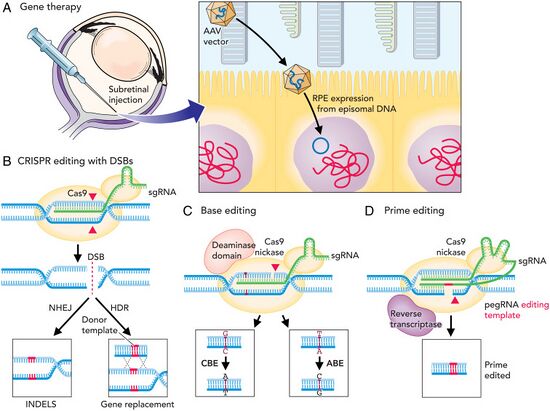
CRISPR-Cas technology has been proposed as a treatment for multiple human diseases, especially those with a genetic cause.[156] Its ability to modify specific DNA sequences makes it a tool with potential to fix disease-causing mutations. Early research in animal models suggest that therapies based on CRISPR technology have potential to treat a wide range of diseases,[157] including cancer,[158] progeria,[159] beta-thalassemia,[160][161][162] sickle cell disease,[162][163] hemophilia,[164] cystic fibrosis,[165] Duchenne's muscular dystrophy,[166] Huntington's disease,[167][168] transthyretin amyloidosis[43] and heart disease.[169] CRISPR has also been used to cure malaria in mosquitos, which could eliminate the vector and the disease in humans.[170] CRISPR may also have applications in tissue engineering and regenerative medicine, such as by creating human blood vessels that lack expression of MHC class II proteins, which often cause transplant rejection.[171]
In addition, clinical trials to cure beta thalassemia and sickle cell disease in human patients using CRISPR-Cas9 technology have shown promising results.[172][173]
Nevertheless, there remains a few limitations of the technology's use in gene therapy: the relatively high frequency of off-target effect, the requirement for a PAM sequence near the target site, p53 mediated apoptosis by CRISPR-induced double-strand breaks and immunogenic toxicity due to the delivery system typically by virus. [174]
Blindness
The CRISPR treatment for LCA10 (the most common variant of Leber Congenital Amaurosis which is the leading cause of inherited childhood blindness) modifies the patient's defective photoreceptor gene.
In March 2020, the first patient volunteer in this US-based study, sponsored by Editas Medicine, was given a low-dose of the treatment to test for safety.
In June 2021, enrollment began for a high-dose adult and pediatric cohort of 4 patient volunteers each. Dosing of the new cohorts is expected to be completed by July 2022.[175] In November 2022, Editas reported that 20% of the patients treated had significant improvements, but also announced that the resulting target population was too small to support continued independent development.[176]
Cancer
CRISPR has also found many applications in developing cell-based immunotherapies.[177] The first clinical trial involving CRISPR started in 2016. It involved taking immune cells from people with lung cancer, using CRISPR to edit out the gene expressed PD-1, then administering the altered cells back to the same person. 20 other trials were under way or nearly ready, mostly in China, as of 2017[update].[158]
In 2016, the United States Food and Drug Administration (FDA) approved a clinical trial in which CRISPR would be used to alter T cells extracted from people with different kinds of cancer and then administer those engineered T cells back to the same people.[178]
In November 2020, in mouse animal models, CRISPR was used effectively to treat glioblastoma (fast-growing brain tumor) and metastatic ovarian cancer, as those are two cancers with some of the worst best-case prognosis and are typically diagnosed during their later stages. The treatments have resulted in inhibited tumor growth, and increased survival by 80% for metastatic ovarian cancer and tumor cell apoptosis, inhibited tumor growth by 50%, and improved survival by 30% for glioblastoma.[179]
In October 2021, CRISPR Therapeutics announced results from their ongoing US-based Phase 1 trial for an allogeneic T cell therapy. These cells are sourced from healthy donors and are edited to attack cancer cells and avoid being seen as a threat by the recipient's immune system, and then multiplied into huge batches which can be given to large numbers of recipients.[175]
In December 2022, a 13-year British girl that had been diagnosed with incurable T-Cell Acute Lymphoblastic Leukaemia was cured by doctors at Great Ormond Street Hospital, in the first documented use of therapeutic gene editing for this purpose, after undergoing six months of an experimental treatment, where previous attempts of other treatments failed. The procedure included reprogramming a healthy T-Cell to destroy the cancerous T-Cells to first rid her of Leukaemia, and then rebuilding her immune system from scratch using healthy immune cells.[180] The team used BASE editing and had previously treated a case of acute lymphoblastic leukaemia in 2015 using TALENs.[181]
Diabetes
Type 1 Diabetes is an endocrine disorder which results from a lack of pancreatic beta cells to produce insulin, a vital compound in transporting blood sugar to cells for producing energy. Researchers have been trying to transplant healthy beta cells. CRISPR is used to edit the cells in order to reduce the chance the patient's body will reject the transplant.
In February 2022, a phase 1 trial was conducted in which one patient volunteer received treatment.[175][182]
HIV/AIDS
Human immunodeficiency virus or HIV, is a virus that attacks the body's immune system. While effective treatments exist which can allow patients to live healthy lives, HIV is retroactive meaning that it embeds an inactive version of itself in the human genome. CRISPR can be used to selectively remove the virus from the genome by designing guide RNA to target the retroactive HIV genome. One issue with this approach is that it requires the removal of the HIV genome from almost all cells, which can be difficult to realistically achieve.[175]
Infection
CRISPR-Cas-based "RNA-guided nucleases" can be used to target virulence factors, genes encoding antibiotic resistance, and other medically relevant sequences of interest. This technology thus represents a novel form of antimicrobial therapy and a strategy by which to manipulate bacterial populations.[183][184] Recent studies suggest a correlation between the interfering of the CRISPR-Cas locus and acquisition of antibiotic resistance.[185] This system provides protection of bacteria against invading foreign DNA, such as transposons, bacteriophages, and plasmids. This system was shown to be a strong selective pressure for the acquisition of antibiotic resistance and virulence factor in bacterial pathogens.[185]
Therapies based on CRISPR–Cas3 gene editing technology delivered by engineered bacteriophages could be used to destroy targeted DNA in pathogens.[186] Cas3 is more destructive than the better known Cas9.[187][188]
Research suggests that CRISPR is an effective way to limit replication of multiple herpesviruses. It was able to eradicate viral DNA in the case of Epstein–Barr virus (EBV). Anti-herpesvirus CRISPRs have promising applications such as removing cancer-causing EBV from tumor cells, helping rid donated organs for immunocompromised patients of viral invaders, or preventing cold sore outbreaks and recurrent eye infections by blocking HSV-1 reactivation. As of August 2016[update], these were awaiting testing.[189]
Initial results in the treatment and cure of HIV have been rather successful, in 2021 9 out of 23 humanized mice treated with a combination of anti-retrovirals and CRISPR/Cas-9 which lead to the virus becoming undetectable, even after the usual rebound period. None of the two treatments alone had such an effect.[190] Clinical trials in humans are to start in 2022.[191][192]
CRISPR may revive the concept of transplanting animal organs into people. Retroviruses present in animal genomes could harm transplant recipients. In 2015, a team eliminated 62 copies of a particular retroviral DNA sequence from the pig genome in a kidney epithelial cell.[193] Researchers recently demonstrated the ability to birth live pig specimens after removing these retroviruses from their genome using CRISPR for the first time.[194]
Genetic anthropology
CRISPR-Cas9 can be used in investigating and identifying the genetic differences of humans to other apes, especially of the brain. For example, by reintroducing archaic gene variants into brain organoids to show an impact on neurogenesis,[195] metaphase length of apical progenitors of the developing neocortex,[196] or by knockout of a gene in embryonic stem cells to identify a genetic regulator that via early cell shape transition drives evolutionary expansion of the human forebrain.[197][198] One study described a major impact of an archaic gene variant on neurodevelopment[199][200] which may be an artefact of a CRISPR side effect,[201][202] as it could not be replicated in a subsequent study.[203]
By technique
Knockdown/activation
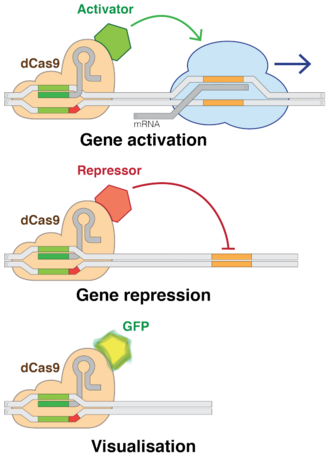
Using "dead" versions of Cas9 (dCas9) eliminates CRISPR's DNA-cutting ability, while preserving its ability to target desirable sequences. Multiple groups added various regulatory factors to dCas9s, enabling them to turn almost any gene on or off or adjust its level of activity.[193] Like RNAi, CRISPR interference (CRISPRi) turns off genes in a reversible fashion by targeting, but not cutting a site. The targeted site is methylated, epigenetically modifying the gene. This modification inhibits transcription. These precisely placed modifications may then be used to regulate the effects on gene expressions and DNA dynamics after the inhibition of certain genome sequences within DNA. Within the past few years, epigenetic marks in different human cells have been closely researched and certain patterns within the marks have been found to correlate with everything ranging from tumor growth to brain activity.[10] Conversely, CRISPR-mediated activation (CRISPRa) promotes gene transcription.[204] Cas9 is an effective way of targeting and silencing specific genes at the DNA level.[205] In bacteria, the presence of Cas9 alone is enough to block transcription. For mammalian applications, a section of protein is added. Its guide RNA targets regulatory DNA sequences called promoters that immediately precede the target gene.[206]
Cas9 was used to carry synthetic transcription factors that activated specific human genes. The technique achieved a strong effect by targeting multiple CRISPR constructs to slightly different locations on the gene's promoter.[206]
RNA editing
In 2016, researchers demonstrated that CRISPR from an ordinary mouth bacterium could be used to edit RNA. The researchers searched databases containing hundreds of millions of genetic sequences for those that resembled CRISPR genes. They considered the fusobacterium Leptotrichia shahii. It had a group of genes that resembled CRISPR genes, but with important differences. When the researchers equipped other bacteria with these genes, which they called C2c2, they found that the organisms gained a novel defense.[207] C2c2 has later been renamed to Cas13a to fit the standard nomenclature for Cas genes.[208]
Many viruses encode their genetic information in RNA rather than DNA that they repurpose to make new viruses. HIV and poliovirus are such viruses. Bacteria with Cas13 make molecules that can dismember RNA, destroying the virus. Tailoring these genes opened any RNA molecule to editing.[207]
CRISPR-Cas systems can also be employed for editing of micro-RNA and long-noncoding RNA genes in plants.[209]
Gene drive
Gene drives may provide a powerful tool to restore balance of ecosystems by eliminating invasive species. Concerns regarding efficacy, unintended consequences in the target species as well as non-target species have been raised particularly in the potential for accidental release from laboratories into the wild. Scientists have proposed several safeguards for ensuring the containment of experimental gene drives including molecular, reproductive, and ecological.[210] Many recommend that immunization and reversal drives be developed in tandem with gene drives in order to overwrite their effects if necessary.[211] There remains consensus that long-term effects must be studied more thoroughly particularly in the potential for ecological disruption that cannot be corrected with reversal drives.[212]
In vitro genetic depletion
Unenriched sequencing libraries often have abundant undesired sequences. Cas9 can specifically deplete the undesired sequences with double strand breakage with up to 99% efficiency and without significant off-target effects as seen with restriction enzymes. Treatment with Cas9 can deplete abundant rRNA while increasing pathogen sensitivity in RNA-seq libraries.[213]
CRISPR-directed integrases
Combination of CRISPR-Cas9 with integrases enabled a technique for large edits without problematic double-stranded breaks, as demonstrated with PASTE in 2022. The researchers reported it could be used to deliver genes as long as 36,000 DNA base pairs to several types of human cells and thereby potentially for treating diseases caused by a large number of mutations.[214][215]
Prime editing
Prime editing[216] (or base editing) is a CRISPR refinement to accurately insert or delete sections of DNA. The CRISPR edits are not always perfect and the cuts can end up in the wrong place. Both issues are a problem for using the technology in medicine.[217] Prime editing does not cut the double-stranded DNA but instead uses the CRISPR targeting apparatus to shuttle an additional enzyme to a desired sequence, where it converts a single nucleotide into another.[218] The new guide, called a pegRNA, contains an RNA template for a new DNA sequence to be added to the genome at the target location. That requires a second protein, attached to Cas9: a reverse transcriptase enzyme, which can make a new DNA strand from the RNA template and insert it at the nicked site.[219] Those three independent pairing events each provide an opportunity to prevent off-target sequences, which significantly increases targeting flexibility and editing precision.[218] Prime editing was developed by researchers at the Broad Institute of MIT and Harvard in Massachusetts.[220] More work is needed to optimize the methods.[220][219]
Society and culture
Human germline modification
As of March 2015, multiple groups had announced ongoing research with the intention of laying the foundations for applying CRISPR to human embryos for human germline engineering, including labs in the US, China, and the UK, as well as US biotechnology company OvaScience.[221] Scientists, including a CRISPR co-discoverer, urged a worldwide moratorium on applying CRISPR to the human germline, especially for clinical use. They said "scientists should avoid even attempting, in lax jurisdictions, germline genome modification for clinical application in humans" until the full implications "are discussed among scientific and governmental organizations".[222][223] These scientists support further low-level research on CRISPR and do not see CRISPR as developed enough for any clinical use in making heritable changes to humans.[224]
In April 2015, Chinese scientists reported results of an attempt to alter the DNA of non-viable human embryos using CRISPR to correct a mutation that causes beta thalassemia, a lethal heritable disorder.[225][226] The study had previously been rejected by both Nature and Science in part because of ethical concerns.[227] The experiments resulted in successfully changing only some of the intended genes, and had off-target effects on other genes. The researchers stated that CRISPR is not ready for clinical application in reproductive medicine.[227] In April 2016, Chinese scientists were reported to have made a second unsuccessful attempt to alter the DNA of non-viable human embryos using CRISPR – this time to alter the CCR5 gene to make the embryo resistant to HIV infection.[228]
In December 2015, an International Summit on Human Gene Editing took place in Washington under the guidance of David Baltimore. Members of national scientific academies of the US, UK, and China discussed the ethics of germline modification. They agreed to support basic and clinical research under certain legal and ethical guidelines. A specific distinction was made between somatic cells, where the effects of edits are limited to a single individual, and germline cells, where genome changes can be inherited by descendants. Heritable modifications could have unintended and far-reaching consequences for human evolution, genetically (e.g. gene–environment interactions) and culturally (e.g. social Darwinism). Altering of gametocytes and embryos to generate heritable changes in humans was defined to be irresponsible. The group agreed to initiate an international forum to address such concerns and harmonize regulations across countries.[229]
In February 2017, the United States National Academies of Sciences, Engineering, and Medicine (NASEM) Committee on Human Gene Editing published a report reviewing ethical, legal, and scientific concerns of genomic engineering technology. The conclusion of the report stated that heritable genome editing is impermissible now but could be justified for certain medical conditions; however, they did not justify the usage of CRISPR for enhancement.[230]
In November 2018, Jiankui He announced that he had edited two human embryos to attempt to disable the gene for CCR5, which codes for a receptor that HIV uses to enter cells. He said that twin girls, Lulu and Nana, had been born a few weeks earlier. He said that the girls still carried functional copies of CCR5 along with disabled CCR5 (mosaicism) and were still vulnerable to HIV. The work was widely condemned as unethical, dangerous, and premature.[231] An international group of scientists called for a global moratorium on genetically editing human embryos.[232]
Policy barriers to genetic engineering
Policy regulations for the CRISPR-Cas9 system vary around the globe. In February 2016, British scientists were given permission by regulators to genetically modify human embryos by using CRISPR-Cas9 and related techniques. However, researchers were forbidden from implanting the embryos and the embryos were to be destroyed after seven days.[233]
The US has an elaborate, interdepartmental regulatory system to evaluate new genetically modified foods and crops. For example, the Agriculture Risk Protection Act of 2000 gives the United States Department of Agriculture the authority to oversee the detection, control, eradication, suppression, prevention, or retardation of the spread of plant pests or noxious weeds to protect the agriculture, environment, and economy of the US. The act regulates any genetically modified organism that utilizes the genome of a predefined "plant pest" or any plant not previously categorized.[234] In 2015, Yinong Yang successfully deactivated 16 specific genes in the white button mushroom to make them non-browning. Since he had not added any foreign-species (transgenic) DNA to his organism, the mushroom could not be regulated by the USDA under Section 340.2.[235] Yang's white button mushroom was the first organism genetically modified with the CRISPR-Cas9 protein system to pass US regulation.[236]
In 2016, the USDA sponsored a committee to consider future regulatory policy for upcoming genetic modification techniques. With the help of the US National Academies of Sciences, Engineering, and Medicine, special interests groups met on April 15 to contemplate the possible advancements in genetic engineering within the next five years and any new regulations that might be needed as a result.[237] In 2017, the Food and Drug Administration proposed a rule that would classify genetic engineering modifications to animals as "animal drugs", subjecting them to strict regulation if offered for sale and reducing the ability for individuals and small businesses to make them profitable.[238][239]
In China, where social conditions sharply contrast with those of the West, genetic diseases carry a heavy stigma.[240] This leaves China with fewer policy barriers to the use of this technology.[241][242]
Recognition
In 2012 and 2013, CRISPR was a runner-up in Science Magazine's Breakthrough of the Year award. In 2015, it was the winner of that award.[193] CRISPR was named as one of MIT Technology Review's 10 breakthrough technologies in 2014 and 2016.[243][244] In 2016, Jennifer Doudna and Emmanuelle Charpentier, along with Rudolph Barrangou, Philippe Horvath, and Feng Zhang won the Gairdner International award. In 2017, Doudna and Charpentier were awarded the Japan Prize in Tokyo, Japan for their revolutionary invention of CRISPR-Cas9. In 2016, Charpentier, Doudna, and Zhang won the Tang Prize in Biopharmaceutical Science.[245] In 2020, Charpentier and Doudna were awarded the Nobel Prize in Chemistry, the first such prize for an all-female team, "for the development of a method for genome editing."[246]
See also
- CRISPR/Cas Tools
- The CRISPR Journal
- Eugenics
- DRACO
- Zinc finger
- Gene knockout
- Genetics
- Glossary of genetics
- Human Nature (2019 documentary film)
- LEAPER gene editing
- Make People Better (2022 documentary)
- RNAi
- SiRNA
- Surveyor nuclease assay
- Synthetic biology
References
- ↑ 1.0 1.1 1.2 Bak RO, Gomez-Ospina N, Porteus MH (August 2018). "Gene Editing on Center Stage". Trends in Genetics. 34 (8): 600–611. doi:10.1016/j.tig.2018.05.004. PMID 29908711. S2CID 49269023.
- ↑ "The Nobel Prize in Chemistry 2020". The Nobel Prize. Archived from the original on 2020-12-11. Retrieved 2020-12-10.
- ↑ Cohen J (October 7, 2020). "CRISPR, the revolutionary genetic "scissors," honored by Chemistry Nobel". Science. doi:10.1126/science.abf0540. S2CID 225116732. Archived from the original on August 29, 2023. Retrieved November 21, 2023.
- ↑ Cohen J (2018-06-04). "With prestigious prize, an overshadowed CRISPR researcher wins the spotlight". Science | AAAS. Archived from the original on 2023-11-25. Retrieved 2020-05-02.
- ↑ 5.0 5.1 Owens R (8 October 2020). "Nobel prize: who gets left out?". The Conversation. Archived from the original on 13 December 2021. Retrieved 13 December 2021.
- ↑ "Lithuanian scientists not awarded Nobel prize despite discovering same technology". LRT.LT. 8 October 2020. Archived from the original on 11 February 2021. Retrieved 21 November 2023.
- ↑ Šikšnys V (2018-06-16). "Imam genų žirkles, iškerpam klaidą, ligos nelieka". Laisvės TV / Freedom TV. 12:22 minutes in. LaisvėsTV. Archived from the original on 2022-01-29. <...>Tai mes tą savo straipsnį išsiuntėm į redakciją pirmieji, bet laimės ten daug nebuvo. Viena redakcija pasakė, kad mes net recenzentam nesiųsim. Nusiuntėm į kitą redakciją – tai jis (straipsnis) pragulėjo kažkur ant redaktoriaus stalo labai ilgai. Na ir taip galų gale išsiuntėm į trečią žurnalą ir trečias žurnalas po kelių mėnesių jį išspausdino. Bet, aišku, Berklio universiteto mokslininkams sekėsi geriau – jie išsiuntė straipsnį į žurnalą Science – jį priėmė ir išspausdino per 2 savaites. Nors iš tikro jie tą straispnį išsiuntė pora mėnesių vėliau nei mes. Retrieved 2018-06-30.
<...> Well, we were who had sent the article first, but had not much of luck.
- ↑ Zhang JH, Pandey M, Kahler JF, Loshakov A, Harris B, Dagur PK, et al. (November 2014). "Improving the specificity and efficacy of CRISPR/CAS9 and gRNA through target specific DNA reporter". Journal of Biotechnology. 189: 1–8. doi:10.1016/j.jbiotec.2014.08.033. PMC 4252756. PMID 25193712.
- ↑ Vakulskas CA, Dever DP, Rettig GR, Turk R, Jacobi AM, Collingwood MA, et al. (August 2018). "A high-fidelity Cas9 mutant delivered as a ribonucleoprotein complex enables efficient gene editing in human hematopoietic stem and progenitor cells". Nature Medicine. 24 (8): 1216–1224. doi:10.1038/s41591-018-0137-0. PMC 6107069. PMID 30082871.
- ↑ 10.0 10.1 10.2 Ledford H (March 2016). "CRISPR: gene editing is just the beginning". Nature. 531 (7593): 156–159. Bibcode:2016Natur.531..156L. doi:10.1038/531156a. PMID 26961639.
- ↑ Travis J (17 December 2015). "Breakthrough of the Year: CRISPR makes the cut". Science Magazine. American Association for the Advancement of Science. Archived from the original on 21 July 2017. Retrieved 21 November 2023.
- ↑ Ledford H (June 2015). "CRISPR, the disruptor". Nature. 522 (7554): 20–24. Bibcode:2015Natur.522...20L. doi:10.1038/522020a. PMID 26040877.
- ↑ 13.0 13.1 "Casgevy: UK approves gene-editing drug for sickle cell". BBC News. 16 November 2023. Archived from the original on 16 November 2023. Retrieved 16 November 2023.
- ↑ 14.0 14.1 "MHRA authorises world-first gene therapy that aims to cure sickle-cell disease and transfusion-dependent β-thalassemia". Gov.uk. 16 November 2023. Archived from the original on 25 November 2023. Retrieved 16 November 2023.
- ↑ Young S (11 February 2014). "CRISPR and Other Genome Editing Tools Boost Medical Research and Gene Therapy's Reach". MIT Technology Review. Archived from the original on 2016-01-12. Retrieved 2014-04-13.
- ↑ 16.0 16.1 Heidenreich M, Zhang F (January 2016). "Applications of CRISPR-Cas systems in neuroscience". Nature Reviews. Neuroscience. 17 (1): 36–44. doi:10.1038/nrn.2015.2. PMC 4899966. PMID 26656253.
- ↑ Barrangou R, Doudna JA (September 2016). "Applications of CRISPR technologies in research and beyond". Nature Biotechnology. 34 (9): 933–941. doi:10.1038/nbt.3659. PMID 27606440. S2CID 21543486.
- ↑ Cox DB, Platt RJ, Zhang F (February 2015). "Therapeutic genome editing: prospects and challenges". Nature Medicine. 21 (2): 121–131. doi:10.1038/nm.3793. PMC 4492683. PMID 25654603.
- ↑ Pollack A (May 11, 2015). "Jennifer Doudna, a Pioneer Who Helped Simplify Genome Editing". New York Times. Archived from the original on January 2, 2022. Retrieved October 8, 2020.
- ↑ Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (August 2012). "A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity". Science. 337 (6096): 816–821. Bibcode:2012Sci...337..816J. doi:10.1126/science.1225829. PMC 6286148. PMID 22745249.
- ↑ "CRISPR Madness". GEN. 2013-11-08. Archived from the original on 2016-10-01. Retrieved 2023-11-21.
- ↑ Staff (1 April 2015). "News: Products & Services". Genetic Engineering & Biotechnology News (Paper). 35 (7): 8. doi:10.1089/gen.35.21.05.
- ↑ "Who Owns the Biggest Biotech Discovery of the Century? There's a bitter fight over the patents for CRISPR, a breakthrough new form of DNA editing". MIT Technology Review. Archived from the original on 29 July 2020. Retrieved 25 February 2015.
- ↑ Fye S. "Genetic Rough Draft: Editas and CRISPR". The Atlas Business Journal. Archived from the original on 26 January 2016. Retrieved 19 January 2016.
- ↑ "CRISPR-Cas systems and methods for altering expression of gene products". Google Patents. Archived from the original on 2023-11-25. Retrieved 2023-11-21.
- ↑ Shaffer C (April 2022). "Broad defeats Berkeley CRISPR patent". Nature Biotechnology. 40 (4): 445. doi:10.1038/d41587-022-00004-2. PMID 35288688. S2CID 247453528.
- ↑ "CRISPR patents to go on trial". Nature Biotechnology. 34 (2): 121. February 2016. doi:10.1038/nbt0216-121a. PMID 26849500. S2CID 205265912.
- ↑ Pollack A (15 February 2017). "Harvard and M.I.T. Scientists Win Gene-Editing Patent Fight". The New York Times. Archived from the original on 25 November 2023. Retrieved 21 November 2023.
- ↑ Akst J (February 15, 2017). "Broad Wins CRISPR Patent Interference Case". The Scientist Magazine. Archived from the original on November 11, 2020. Retrieved November 21, 2023.
- ↑ Noonan KE (February 16, 2017). "PTAB Decides CRISPR Interference in Favor of Broad Institute – Their Reasoning". Patent Docs. Archived from the original on June 7, 2023. Retrieved November 21, 2023.
- ↑ Potenza A (April 13, 2017). "UC Berkeley challenges decision that CRISPR patents belong to Broad Institute". The Verge. Archived from the original on 22 September 2017. Retrieved 22 September 2017.
- ↑ Buhr S (July 26, 2017). "The CRISPR patent battle is back on as UC Berkeley files an appeal". TechCrunch. Archived from the original on 22 September 2017. Retrieved 22 September 2017.
- ↑ Westman N (March 1, 2022). "UC Berkeley loses CRISPR patent case". The Verge. Archived from the original on April 30, 2022. Retrieved March 6, 2022.
- ↑ 34.0 34.1 Philippidis A (August 7, 2017). "MilliporeSigma to Be Granted European Patent for CRISPR Technology". Genetic Engineering & Biotechology News. Archived from the original on 23 September 2017. Retrieved 22 September 2017.
- ↑ Akst J (March 24, 2017). "UC Berkeley Receives CRISPR Patent in Europe". The Scientist. Archived from the original on 25 November 2023. Retrieved 22 September 2017.
- ↑ Cohen J (4 August 2017). "CRISPR patent battle in Europe takes a 'wild' twist with surprising player". Science. doi:10.1126/science.aan7211. Archived from the original on 6 April 2023. Retrieved 21 November 2023.
- ↑ "Top EU court: GMO rules cover plant gene editing technique". Retuers. 25 July 2018. Archived from the original on 7 April 2023. Retrieved 21 November 2023.
- ↑ AFP (7 February 2020). "US Trial Shows 3 Cancer Patients Had Their Genomes Altered Safely by CRISPR". ScienceAlert. Archived from the original on 2020-02-08. Retrieved 2020-02-09.
- ↑ Chamary JV. "These Scientists Deserved A Nobel Prize, But Didn't Discover Crispr". Forbes. Archived from the original on 2020-10-08. Retrieved 2020-07-10.
- ↑ Fischman J. "Nobel Prize in Chemistry Goes to Discovery of 'Genetic Scissors' Called CRISPR/Cas9". Scientific American. Archived from the original on 2021-03-05. Retrieved 2021-03-24.
- ↑ "Two women share chemistry Nobel in historic win for 'genetic scissors'". BBC News. 2020-10-07. Archived from the original on 2020-12-11. Retrieved 2020-12-06.
- ↑ Kaiser J (26 June 2021). "CRISPR injected into the blood treats a genetic disease for first time". Science | AAAS. Archived from the original on 29 September 2021. Retrieved 11 July 2021.
- ↑ 43.0 43.1 Gillmore JD, Gane E, Taubel J, Kao J, Fontana M, Maitland ML, et al. (August 2021). "CRISPR-Cas9 In Vivo Gene Editing for Transthyretin Amyloidosis". The New England Journal of Medicine. 385 (6): 493–502. doi:10.1056/NEJMoa2107454. PMID 34215024. S2CID 235722446.
- ↑ Boonstra E, de Kleijn R, Colzato LS, Alkemade A, Forstmann BU, Nieuwenhuis S (6 October 2015). "Neurotransmitters as food supplements: the effects of GABA on brain and behavior". Frontiers in Psychology. 6: 1520. doi:10.3389/fpsyg.2015.01520. PMC 4594160. PMID 26500584.
- ↑ "Tomato In Japan Is First CRISPR-Edited Food In The World To Go On Sale". IFLScience. Archived from the original on 25 October 2021. Retrieved 18 October 2021.
- ↑ Wang T, Zhang H, Zhu H (15 June 2019). "CRISPR technology is revolutionizing the improvement of tomato and other fruit crops". Horticulture Research. 6 (1): 77. doi:10.1038/s41438-019-0159-x. PMC 6570646. PMID 31240102.
- ↑ "Japan embraces CRISPR-edited fish". Nature Biotechnology. 40 (1): 10. January 2022. doi:10.1038/s41587-021-01197-8. PMID 34969964. S2CID 245593283.
- ↑ "Startup hopes genome-edited pufferfish will be a hit in 2022". The Japan Times. 5 January 2022. Archived from the original on 17 January 2022. Retrieved 17 January 2022.
- ↑ "Gene-edited sea bream set for sale in Japan". thefishsite.com. Archived from the original on 2023-04-05. Retrieved 2023-11-21.
- ↑ Götz L, Svanidze M, Tissier A, Brand A (January 2022). "Consumers' Willingness to Buy CRISPR Gene-Edited Tomatoes: Evidence from a Choice Experiment Case Study in Germany". Sustainability. 14 (2): 971. doi:10.3390/su14020971.
- ↑ "Are Consumers Willing to Buy CRISPR Tomatoes?". Crop Biotech Update. Archived from the original on 2022-02-16. Retrieved 2022-02-21.
- ↑ Whitford E (2021-05-28). "UC Berkeley Will Auction NFTs for 2 Nobel Prize Patents". Inside Higher Ed. Archived from the original on 2023-01-24. Retrieved 2023-02-21.
- ↑ Sestino A, Guido G, Peluso AM (2022). Non-Fungible Tokens (NFTs). Examining the Impact on Consumers and Marketing Strategies. p. 28. doi:10.1007/978-3-031-07203-1. ISBN 978-3-031-07202-4. S2CID 250238540.
- ↑ Chang K (2021-05-27). "You Can Buy a Piece of a Nobel Prize-Winning Discovery". New York Times. Archived from the original on 2023-03-08. Retrieved 2023-02-21.
- ↑ Trautman LJ (2022). "Virtual Art and Non-Fungible Tokens" (PDF). Hofstra Law Review. 50 (361): 369 f. doi:10.2139/ssrn.3814087. S2CID 234830426. Archived (PDF) from the original on 2023-11-25. Retrieved 2023-11-21.
- ↑ Jones N (2021-06-18). "How scientists are embracing NFTs". Nature. 594 (7864): 482. Bibcode:2021Natur.594..481J. doi:10.1038/d41586-021-01642-3. PMID 34145410. S2CID 235481285.
- ↑ 57.0 57.1 57.2 57.3 57.4 Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F (November 2013). "Genome engineering using the CRISPR-Cas9 system". Nature Protocols. 8 (11): 2281–2308. doi:10.1038/nprot.2013.143. hdl:1721.1/102943. PMC 3969860. PMID 24157548.
- ↑ Ly J (2013). Discovering Genes Responsible for Kidney Diseases (Ph.D.). University of Toronto. Retrieved 26 December 2016.
- ↑ Mohr SE, Hu Y, Ewen-Campen B, Housden BE, Viswanatha R, Perrimon N (September 2016). "CRISPR guide RNA design for research applications". The FEBS Journal. 283 (17): 3232–3238. doi:10.1111/febs.13777. PMC 5014588. PMID 27276584.
- ↑ Brazelton VA, Zarecor S, Wright DA, Wang Y, Liu J, Chen K, et al. (2015). "A quick guide to CRISPR sgRNA design tools". GM Crops & Food. 6 (4): 266–276. doi:10.1080/21645698.2015.1137690. PMC 5033207. PMID 26745836.
- ↑ "Researchers establish new viable CRISPR-Cas12b system for plant genome engineering". phys.org. Archived from the original on 6 April 2020. Retrieved 6 April 2020.
- ↑ Ming M, Ren Q, Pan C, He Y, Zhang Y, Liu S, et al. (March 2020). "CRISPR-Cas12b enables efficient plant genome engineering". Nature Plants. 6 (3): 202–208. doi:10.1038/s41477-020-0614-6. PMID 32170285. S2CID 212643374.
- ↑ Cox DB, Gootenberg JS, Abudayyeh OO, Franklin B, Kellner MJ, Joung J, Zhang F (November 2017). "RNA editing with CRISPR-Cas13". Science. 358 (6366): 1019–1027. Bibcode:2017Sci...358.1019C. doi:10.1126/science.aaq0180. PMC 5793859. PMID 29070703.
- ↑ "CRISPR-Cas3 innovation holds promise for disease cures, advancing science". Cornell Chronicle. Archived from the original on 24 October 2021. Retrieved 24 October 2021.
- ↑ Dolan AE, Hou Z, Xiao Y, Gramelspacher MJ, Heo J, Howden SE, et al. (June 2019). "Introducing a Spectrum of Long-Range Genomic Deletions in Human Embryonic Stem Cells Using Type I CRISPR-Cas". Molecular Cell. 74 (5): 936–950.e5. doi:10.1016/j.molcel.2019.03.014. PMC 6555677. PMID 30975459.
- ↑ Liu Z, Dong H, Cui Y, Cong L, Zhang D (September 2020). "Application of different types of CRISPR/Cas-based systems in bacteria". Microbial Cell Factories. 19 (1): 172. doi:10.1186/s12934-020-01431-z. PMC 7470686. PMID 32883277.
- ↑ "Researchers develop an engineered 'mini' CRISPR genome editing system". phys.org. Archived from the original on 18 October 2021. Retrieved 18 October 2021.
- ↑ Xu X, Chemparathy A, Zeng L, Kempton HR, Shang S, Nakamura M, Qi LS (October 2021). "Engineered miniature CRISPR-Cas system for mammalian genome regulation and editing". Molecular Cell. 81 (20): 4333–4345.e4. doi:10.1016/j.molcel.2021.08.008. PMID 34480847. S2CID 237417317.
- ↑ Bravo JP, Liu MS, Hibshman GN, Dangerfield TL, Jung K, McCool RS, et al. (March 2022). "Structural basis for mismatch surveillance by CRISPR-Cas9". Nature. 603 (7900): 343–347. Bibcode:2022Natur.603..343B. doi:10.1038/s41586-022-04470-1. PMC 8907077. PMID 35236982.
- ↑ "Protein tweak makes CRISPR gene editing 4,000 times less error-prone". New Atlas. 2022-03-04. Archived from the original on 2022-03-06. Retrieved 2022-03-07.
- ↑ Kato K, Zhou W, Okazaki S, Isayama Y, Nishizawa T, Gootenberg JS, et al. (June 2022). "Structure and engineering of the type III-E CRISPR-Cas7-11 effector complex". Cell. 185 (13): 2324–2337.e16. doi:10.1016/j.cell.2022.05.003. PMID 35643083. S2CID 249103058.
- Lay summary: Williams S. "Neuroscientists expand CRISPR toolkit with new, compact Cas7-11 enzyme". Massachusetts Institute of Technology. Archived from the original on 22 June 2022. Retrieved 22 June 2022.
- ↑ "'Softer' form of CRISPR may edit genes more accurately". New Scientist. Archived from the original on 21 August 2022. Retrieved 21 August 2022.
- ↑ Roy S, Juste SS, Sneider M, Auradkar A, Klanseck C, Li Z, et al. (July 2022). "Cas9/Nickase-induced allelic conversion by homologous chromosome-templated repair in Drosophila somatic cells". Science Advances. 8 (26): eabo0721. Bibcode:2022SciA....8O.721R. doi:10.1126/sciadv.abo0721. PMID 35776792.
- ↑ Horvath P, Barrangou R (January 2010). "CRISPR/Cas, the immune system of bacteria and archaea". Science. 327 (5962): 167–170. Bibcode:2010Sci...327..167H. doi:10.1126/science.1179555. PMID 20056882. S2CID 17960960.
- ↑ Bialk P, Rivera-Torres N, Strouse B, Kmiec EB (2015-06-08). "Regulation of Gene Editing Activity Directed by Single-Stranded Oligonucleotides and CRISPR/Cas9 Systems". PLOS ONE. 10 (6): e0129308. Bibcode:2015PLoSO..1029308B. doi:10.1371/journal.pone.0129308. PMC 4459703. PMID 26053390.
- ↑ Sander JD, Joung JK (April 2014). "CRISPR-Cas systems for editing, regulating and targeting genomes". Nature Biotechnology. 32 (4): 347–355. doi:10.1038/nbt.2842. PMC 4022601. PMID 24584096.
- ↑ Riesenberg, Stephan; Kanis, Philipp; Macak, Dominik; Wollny, Damian; Düsterhöft, Dorothee; Kowalewski, Johannes; Helmbrecht, Nelly; Maricic, Tomislav; Pääbo, Svante (2023-07-20). "Efficient high-precision homology-directed repair-dependent genome editing by HDRobust". Nature Methods. 20 (9): 1388–1399. doi:10.1038/s41592-023-01949-1. ISSN 1548-7091. PMC 10482697. PMID 37474806.
{{cite journal}}: Check|pmc=value (help) - ↑ Lino CA, Harper JC, Carney JP, Timlin JA (November 2018). "Delivering CRISPR: a review of the challenges and approaches". Drug Delivery. 25 (1): 1234–1257. doi:10.1080/10717544.2018.1474964. PMC 6058482. PMID 29801422.
- ↑ Li L, Hu S, Chen X (July 2018). "Non-viral delivery systems for CRISPR/Cas9-based genome editing: Challenges and opportunities". Biomaterials. 171: 207–218. doi:10.1016/j.biomaterials.2018.04.031. PMC 5944364. PMID 29704747.
- ↑ Jain PK, Lo JH, Rananaware S, Downing M, Panda A, Tai M, et al. (November 2019). "Non-viral delivery of CRISPR/Cas9 complex using CRISPR-GPS nanocomplexes". Nanoscale. 11 (44): 21317–21323. doi:10.1039/C9NR01786K. PMC 7709491. PMID 31670340.
- ↑ 81.0 81.1 Yip BH (May 2020). "Recent Advances in CRISPR/Cas9 Delivery Strategies". Biomolecules. 10 (6): 839. doi:10.3390/biom10060839. PMC 7356196. PMID 32486234.
- ↑ Bak RO, Porteus MH (July 2017). "CRISPR-Mediated Integration of Large Gene Cassettes Using AAV Donor Vectors". Cell Reports. 20 (3): 750–756. doi:10.1016/j.celrep.2017.06.064. PMC 5568673. PMID 28723575.
- ↑ Schmidt F, Grimm D (February 2015). "CRISPR genome engineering and viral gene delivery: a case of mutual attraction". Biotechnology Journal. 10 (2): 258–272. doi:10.1002/biot.201400529. PMID 25663455. S2CID 37653318.
- ↑ Waxmonsky N (24 September 2015). "CRISPR 101: Mammalian Expression Systems and Delivery Methods". Archived from the original on 12 June 2018. Retrieved 11 June 2018.
- ↑ Mishra T, Bhardwaj V, Ahuja N, Gadgil P, Ramdas P, Shukla S, Chande A (June 2022). "Improved loss-of-function CRISPR-Cas9 genome editing in human cells concomitant with inhibition of TGF-β signaling". Molecular Therapy. Nucleic Acids. 28: 202–218. doi:10.1016/j.omtn.2022.03.003. PMC 8961078. PMID 35402072.
- ↑ Adli M (May 2018). "The CRISPR tool kit for genome editing and beyond". Nature Communications. 9 (1): 1911. Bibcode:2018NatCo...9.1911A. doi:10.1038/s41467-018-04252-2. PMC 5953931. PMID 29765029.
- ↑ Svitashev S, Schwartz C, Lenderts B, Young JK, Mark Cigan A (November 2016). "Genome editing in maize directed by CRISPR-Cas9 ribonucleoprotein complexes". Nature Communications. 7 (1): 13274. Bibcode:2016NatCo...713274S. doi:10.1038/ncomms13274. PMC 5116081. PMID 27848933.
- ↑ Liang Z, Chen K, Li T, Zhang Y, Wang Y, Zhao Q, et al. (January 2017). "Efficient DNA-free genome editing of bread wheat using CRISPR/Cas9 ribonucleoprotein complexes". Nature Communications. 8 (1): 14261. Bibcode:2017NatCo...814261L. doi:10.1038/ncomms14261. PMC 5253684. PMID 28098143. S2CID 17028472.
- ↑ Cui Z, Tian R, Huang Z, Jin Z, Li L, Liu J, et al. (March 2022). "FrCas9 is a CRISPR/Cas9 system with high editing efficiency and fidelity". Nature Communications. 13 (1): 1425. Bibcode:2022NatCo..13.1425C. doi:10.1038/s41467-022-29089-8. PMC 8931148. PMID 35301321.
- ↑ Kleinstiver BP, Pattanayak V, Prew MS, Tsai SQ, Nguyen NT, Zheng Z, Joung JK (January 2016). "High-fidelity CRISPR-Cas9 nucleases with no detectable genome-wide off-target effects". Nature. 529 (7587): 490–495. Bibcode:2016Natur.529..490K. doi:10.1038/nature16526. PMC 4851738. PMID 26735016.
- ↑ Slaymaker IM, Gao L, Zetsche B, Scott DA, Yan WX, Zhang F (January 2016). "Rationally engineered Cas9 nucleases with improved specificity". Science. 351 (6268): 84–88. Bibcode:2016Sci...351...84S. doi:10.1126/science.aad5227. PMC 4714946. PMID 26628643.
- ↑ Yin H, Song CQ, Suresh S, Kwan SY, Wu Q, Walsh S, et al. (March 2018). "Partial DNA-guided Cas9 enables genome editing with reduced off-target activity". Nature Chemical Biology. 14 (3): 311–316. doi:10.1038/nchembio.2559. PMC 5902734. PMID 29377001.
- ↑ Riesenberg S, Helmbrecht N, Kanis P, Maricic T, Pääbo S (January 2022). "Improved gRNA secondary structures allow editing of target sites resistant to CRISPR-Cas9 cleavage". Nature Communications. 13 (1): 489. Bibcode:2022NatCo..13..489R. doi:10.1038/s41467-022-28137-7. PMC 8789806. PMID 35078986. S2CID 246281892.
- ↑ Fu Y, Sander JD, Reyon D, Cascio VM, Joung JK (March 2014). "Improving CRISPR-Cas nuclease specificity using truncated guide RNAs". Nature Biotechnology. 32 (3): 279–284. doi:10.1038/nbt.2808. PMC 3988262. PMID 24463574.
- ↑ Thean DG, Chu HY, Fong JH, Chan BK, Zhou P, Kwok CC, et al. (April 2022). "Machine learning-coupled combinatorial mutagenesis enables resource-efficient engineering of CRISPR-Cas9 genome editor activities". Nature Communications. 13 (1): 2219. Bibcode:2022NatCo..13.2219T. doi:10.1038/s41467-022-29874-5. PMC 9039034. PMID 35468907.
- ↑ Cho SW, Kim S, Kim Y, Kweon J, Kim HS, Bae S, Kim JS (January 2014). "Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases". Genome Research. 24 (1): 132–141. doi:10.1101/gr.162339.113. PMC 3875854. PMID 24253446.
- ↑ 97.0 97.1 Oakes BL, Nadler DC, Flamholz A, Fellmann C, Staahl BT, Doudna JA, Savage DF (June 2016). "Profiling of engineering hotspots identifies an allosteric CRISPR-Cas9 switch". Nature Biotechnology. 34 (6): 646–651. doi:10.1038/nbt.3528. PMC 4900928. PMID 27136077.
- ↑ Nuñez JK, Harrington LB, Doudna JA (March 2016). "Chemical and Biophysical Modulation of Cas9 for Tunable Genome Engineering". ACS Chemical Biology. 11 (3): 681–688. doi:10.1021/acschembio.5b01019. PMID 26857072.
- ↑ Zhou W, Deiters A (April 2016). "Conditional Control of CRISPR/Cas9 Function". Angewandte Chemie. 55 (18): 5394–5399. doi:10.1002/anie.201511441. PMID 26996256.
- ↑ Polstein LR, Gersbach CA (March 2015). "A light-inducible CRISPR-Cas9 system for control of endogenous gene activation". Nature Chemical Biology. 11 (3): 198–200. doi:10.1038/nchembio.1753. PMC 4412021. PMID 25664691.
- ↑ Nihongaki Y, Yamamoto S, Kawano F, Suzuki H, Sato M (February 2015). "CRISPR-Cas9-based photoactivatable transcription system". Chemistry & Biology. 22 (2): 169–174. doi:10.1016/j.chembiol.2014.12.011. PMID 25619936.
- ↑ Wright AV, Sternberg SH, Taylor DW, Staahl BT, Bardales JA, Kornfeld JE, Doudna JA (March 2015). "Rational design of a split-Cas9 enzyme complex". Proceedings of the National Academy of Sciences of the United States of America. 112 (10): 2984–2989. Bibcode:2015PNAS..112.2984W. doi:10.1073/pnas.1501698112. PMC 4364227. PMID 25713377.
- ↑ Nihongaki Y, Kawano F, Nakajima T, Sato M (July 2015). "Photoactivatable CRISPR-Cas9 for optogenetic genome editing". Nature Biotechnology. 33 (7): 755–760. doi:10.1038/nbt.3245. PMID 26076431. S2CID 205281536.
- ↑ Hemphill J, Borchardt EK, Brown K, Asokan A, Deiters A (May 2015). "Optical Control of CRISPR/Cas9 Gene Editing". Journal of the American Chemical Society. 137 (17): 5642–5645. doi:10.1021/ja512664v. PMC 4919123. PMID 25905628.
- ↑ Jain PK, Ramanan V, Schepers AG, Dalvie NS, Panda A, Fleming HE, Bhatia SN (September 2016). "Development of Light-Activated CRISPR Using Guide RNAs with Photocleavable Protectors". Angewandte Chemie. 55 (40): 12440–12444. doi:10.1002/anie.201606123. PMC 5864249. PMID 27554600.
- ↑ Davis KM, Pattanayak V, Thompson DB, Zuris JA, Liu DR (May 2015). "Small molecule-triggered Cas9 protein with improved genome-editing specificity". Nature Chemical Biology. 11 (5): 316–318. doi:10.1038/nchembio.1793. PMC 4402137. PMID 25848930.
- ↑ Liu KI, Ramli MN, Woo CW, Wang Y, Zhao T, Zhang X, et al. (November 2016). "A chemical-inducible CRISPR-Cas9 system for rapid control of genome editing". Nature Chemical Biology. 12 (11): 980–987. doi:10.1038/nchembio.2179. PMID 27618190. S2CID 33891039.
- ↑ Truong DJ, Kühner K, Kühn R, Werfel S, Engelhardt S, Wurst W, Ortiz O (July 2015). "Development of an intein-mediated split-Cas9 system for gene therapy". Nucleic Acids Research. 43 (13): 6450–6458. doi:10.1093/nar/gkv601. PMC 4513872. PMID 26082496.
- ↑ Zetsche B, Volz SE, Zhang F (February 2015). "A split-Cas9 architecture for inducible genome editing and transcription modulation". Nature Biotechnology. 33 (2): 139–142. doi:10.1038/nbt.3149. PMC 4503468. PMID 25643054.
- ↑ González F, Zhu Z, Shi ZD, Lelli K, Verma N, Li QV, Huangfu D (August 2014). "An iCRISPR platform for rapid, multiplexable, and inducible genome editing in human pluripotent stem cells". Cell Stem Cell. 15 (2): 215–226. doi:10.1016/j.stem.2014.05.018. PMC 4127112. PMID 24931489.
- ↑ Dow LE, Fisher J, O'Rourke KP, Muley A, Kastenhuber ER, Livshits G, et al. (April 2015). "Inducible in vivo genome editing with CRISPR-Cas9". Nature Biotechnology. 33 (4): 390–394. doi:10.1038/nbt.3155. PMC 4390466. PMID 25690852.
- ↑ Yu C, Liu Y, Ma T, Liu K, Xu S, Zhang Y, et al. (February 2015). "Small molecules enhance CRISPR genome editing in pluripotent stem cells". Cell Stem Cell. 16 (2): 142–147. doi:10.1016/j.stem.2015.01.003. PMC 4461869. PMID 25658371.
- ↑ Schimmel, J.; Muñoz-Subirana, N.; Kool, H.; Van Schendel, R.; Van Der Vlies, S.; Kamp, J. A.; De Vrij, F.; Kushner, S. A.; Smith GCM; Boulton, S. J.; Tijsterman, M. (2023). "Modulating mutational outcomes and improving precise gene editing at CRISPR-Cas9-induced breaks by chemical inhibition of end-joining pathways". Cell Reports. 42 (2): 112019. doi:10.1016/j.celrep.2023.112019. PMID 36701230. S2CID 256273893. Archived from the original on 2023-07-03. Retrieved 2023-11-21.
- ↑ Maruyama T, Dougan SK, Truttmann MC, Bilate AM, Ingram JR, Ploegh HL (May 2015). "Increasing the efficiency of precise genome editing with CRISPR-Cas9 by inhibition of nonhomologous end joining". Nature Biotechnology. 33 (5): 538–542. doi:10.1038/nbt.3190. PMC 4618510. PMID 25798939.
- ↑ 115.0 115.1 Ma X, Chen X, Jin Y, Ge W, Wang W, Kong L, et al. (April 2018). "Small molecules promote CRISPR-Cpf1-mediated genome editing in human pluripotent stem cells". Nature Communications. 9 (1): 1303. Bibcode:2018NatCo...9.1303M. doi:10.1038/s41467-018-03760-5. PMC 5880812. PMID 29610531.
- ↑ Nishida K, Arazoe T, Yachie N, Banno S, Kakimoto M, Tabata M, et al. (September 2016). "Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems". Science. 353 (6305): aaf8729. doi:10.1126/science.aaf8729. PMID 27492474. S2CID 5122081.
- ↑ Komor AC, Kim YB, Packer MS, Zuris JA, Liu DR (May 2016). "Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage". Nature. 533 (7603): 420–424. Bibcode:2016Natur.533..420K. doi:10.1038/nature17946. PMC 4873371. PMID 27096365.
- ↑ Gaudelli NM, Komor AC, Rees HA, Packer MS, Badran AH, Bryson DI, Liu DR (November 2017). "Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage". Nature. 551 (7681): 464–471. Bibcode:2017Natur.551..464G. doi:10.1038/nature24644. PMC 5726555. PMID 29160308.
- ↑ Chen L, Park JE, Paa P, Rajakumar PD, Prekop HT, Chew YT, et al. (March 2021). "Programmable C:G to G:C genome editing with CRISPR-Cas9-directed base excision repair proteins". Nature Communications. 12 (1): 1384. Bibcode:2021NatCo..12.1384C. doi:10.1038/s41467-021-21559-9. PMC 7925527. PMID 33654077.
- ↑ 120.0 120.1 120.2 Kurata M, Yamamoto K, Moriarity BS, Kitagawa M, Largaespada DA (February 2018). "CRISPR/Cas9 library screening for drug target discovery". Journal of Human Genetics. 63 (2): 179–186. doi:10.1038/s10038-017-0376-9. PMID 29158600. S2CID 3308058.
- ↑ Gasiunas G, Barrangou R, Horvath P, Siksnys V (September 2012). "Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria". Proceedings of the National Academy of Sciences of the United States of America. 109 (39): E2579–E2586. doi:10.1073/pnas.1208507109. PMC 3465414. PMID 22949671.
- ↑ Satomura A, Nishioka R, Mori H, Sato K, Kuroda K, Ueda M (May 2017). "Precise genome-wide base editing by the CRISPR Nickase system in yeast". Scientific Reports. 7 (1): 2095. Bibcode:2017NatSR...7.2095S. doi:10.1038/s41598-017-02013-7. PMC 5437071. PMID 28522803.
- ↑ Hiranniramol K, Chen Y, Liu W, Wang X (May 2020). "Generalizable sgRNA design for improved CRISPR/Cas9 editing efficiency". Bioinformatics. 36 (9): 2684–2689. doi:10.1093/bioinformatics/btaa041. PMC 7203743. PMID 31971562.
- ↑ 124.0 124.1 124.2 Agrotis A, Ketteler R (2015-09-24). "A new age in functional genomics using CRISPR/Cas9 in arrayed library screening". Frontiers in Genetics. 6: 300. doi:10.3389/fgene.2015.00300. PMC 4585242. PMID 26442115.
- ↑ Yu JS, Yusa K (July 2019). "Genome-wide CRISPR-Cas9 screening in mammalian cells". Methods. 164–165: 29–35. doi:10.1016/j.ymeth.2019.04.015. PMID 31034882. S2CID 140275157.
- ↑ 126.0 126.1 Joung J, Konermann S, Gootenberg JS, Abudayyeh OO, Platt RJ, Brigham MD, et al. (April 2017). "Genome-scale CRISPR-Cas9 knockout and transcriptional activation screening". Nature Protocols. 12 (4): 828–863. doi:10.1038/nprot.2017.016. PMC 5526071. PMID 28333914.
- ↑ "Addgene: Pooled Libraries". www.addgene.org. Archived from the original on 2020-02-04. Retrieved 2020-01-31.
- ↑ McDade JR, Waxmonsky NC, Swanson LE, Fan M (July 2016). "Practical Considerations for Using Pooled Lentiviral CRISPR Libraries". Current Protocols in Molecular Biology. 115 (1): 31.5.1–31.5.13. doi:10.1002/cpmb.8. PMID 27366891. S2CID 5055878.
- ↑ Cheng AW, Wang H, Yang H, Shi L, Katz Y, Theunissen TW, et al. (October 2013). "Multiplexed activation of endogenous genes by CRISPR-on, an RNA-guided transcriptional activator system". Cell Research. 23 (10): 1163–1171. doi:10.1038/cr.2013.122. PMC 3790238. PMID 23979020.
- ↑ Gilbert LA, Horlbeck MA, Adamson B, Villalta JE, Chen Y, Whitehead EH, et al. (October 2014). "Genome-Scale CRISPR-Mediated Control of Gene Repression and Activation". Cell. 159 (3): 647–661. doi:10.1016/j.cell.2014.09.029. PMC 4253859. PMID 25307932.
- ↑ 131.0 131.1 Dow LE (October 2015). "Modeling Disease In Vivo With CRISPR/Cas9". Trends in Molecular Medicine. 21 (10): 609–621. doi:10.1016/j.molmed.2015.07.006. PMC 4592741. PMID 26432018.
- ↑ Doudna J, Mali P (2016). CRISPR-Cas: A Laboratory Manual. Cold Spring Harbor, New York. ISBN 978-1-62182-130-4. OCLC 922914104.
- ↑ Zuo E, Huo X, Yao X, Hu X, Sun Y, Yin J, et al. (November 2017). "CRISPR/Cas9-mediated targeted chromosome elimination". Genome Biology. 18 (1): 224. doi:10.1186/s13059-017-1354-4. PMC 5701507. PMID 29178945.
- "CRISPR Used to Eliminate Targeted Chromosomes in New Study". Genome Web. Nov 27, 2017. Archived from the original on April 7, 2023. Retrieved November 21, 2023.
- ↑ Javed MR, Sadaf M, Ahmed T, Jamil A, Nawaz M, Abbas H, Ijaz A (December 2018). "CRISPR-Cas System: History and Prospects as a Genome Editing Tool in Microorganisms". review. Current Microbiology. 75 (12): 1675–1683. doi:10.1007/s00284-018-1547-4. PMID 30078067. S2CID 51920661.
- ↑ DiCarlo JE, Norville JE, Mali P, Rios X, Aach J, Church GM (April 2013). "Genome engineering in Saccharomyces cerevisiae using CRISPR-Cas systems". Nucleic Acids Res. 41 (7): 4336–43. doi:10.1093/nar/gkt135. PMC 3627607. PMID 23460208.
- ↑ Giersch RM, Finnigan GC (December 2017). "Yeast Still a Beast: Diverse Applications of CRISPR/Cas Editing Technology in S. cerevisiae". The Yale Journal of Biology and Medicine. 90 (4): 643–651. PMC 5733842. PMID 29259528.
- ↑ Dhamad AE, Lessner DJ (October 2020). Atomi H (ed.). "A CRISPRi-dCas9 System for Archaea and Its Use To Examine Gene Function during Nitrogen Fixation by Methanosarcina acetivorans". Applied and Environmental Microbiology. 86 (21): e01402–20. Bibcode:2020ApEnM..86E1402D. doi:10.1128/AEM.01402-20. PMC 7580536. PMID 32826220.
- ↑ Raschmanová H, Weninger A, Glieder A, Kovar K, Vogl T (2018). "Implementing CRISPR-Cas technologies in conventional and non-conventional yeasts: Current state and future prospects". review. Biotechnology Advances. 36 (3): 641–665. doi:10.1016/j.biotechadv.2018.01.006. PMID 29331410.
- ↑ 139.0 139.1 Ma D, Liu F (December 2015). "Genome Editing and Its Applications in Model Organisms". review. Genomics, Proteomics & Bioinformatics. 13 (6): 336–344. doi:10.1016/j.gpb.2015.12.001. PMC 4747648. PMID 26762955.
- ↑ Khurshid H, Jan SA, Shinwari ZK, Jamal M, Shah SH (2018). "An Era of CRISPR/ Cas9 Mediated Plant Genome Editing". review. Current Issues in Molecular Biology. 26: 47–54. doi:10.21775/cimb.026.047. PMID 28879855.
- ↑ Simone BW, Martínez-Gálvez G, WareJoncas Z, Ekker SC (November 2018). "Fishing for understanding: Unlocking the zebrafish gene editor's toolbox". review. Methods. 150: 3–10. doi:10.1016/j.ymeth.2018.07.012. PMC 6590056. PMID 30076892.
- ↑ Singh P, Schimenti JC, Bolcun-Filas E (January 2015). "A mouse geneticist's practical guide to CRISPR applications". review. Genetics. 199 (1): 1–15. doi:10.1534/genetics.114.169771. PMC 4286675. PMID 25271304.
- ↑ Soni D, Wang DM, Regmi SC, Mittal M, Vogel SM, Schlüter D, Tiruppathi C (May 2018). "Deubiquitinase function of A20 maintains and repairs endothelial barrier after lung vascular injury". Cell Death Discovery. 4 (60): 60. doi:10.1038/s41420-018-0056-3. PMC 5955943. PMID 29796309.
- ↑ Birling MC, Herault Y, Pavlovic G (August 2017). "Modeling human disease in rodents by CRISPR/Cas9 genome editing". Mammalian Genome. 28 (7–8): 291–301. doi:10.1007/s00335-017-9703-x. PMC 5569124. PMID 28677007.
- ↑ Gao X, Tao Y, Lamas V, Huang M, Yeh WH, Pan B, et al. (January 2018). "Treatment of autosomal dominant hearing loss by in vivo delivery of genome editing agents". Nature. 553 (7687): 217–221. Bibcode:2018Natur.553..217G. doi:10.1038/nature25164. PMC 5784267. PMID 29258297.
- ↑ Kadam US, Shelake RM, Chavhan RL, Suprasanna P (October 2018). "Concerns regarding 'off-target' activity of genome editing endonucleases". review. Plant Physiology and Biochemistry. 131: 22–30. doi:10.1016/j.plaphy.2018.03.027. PMID 29653762. S2CID 4846191.
- ↑ Kimberland ML, Hou W, Alfonso-Pecchio A, Wilson S, Rao Y, Zhang S, Lu Q (October 2018). "Strategies for controlling CRISPR/Cas9 off-target effects and biological variations in mammalian genome editing experiments". review. Journal of Biotechnology. 284: 91–101. doi:10.1016/j.jbiotec.2018.08.007. PMID 30142414. S2CID 52078796.
- ↑ van Erp PB, Bloomer G, Wilkinson R, Wiedenheft B (June 2015). "The history and market impact of CRISPR RNA-guided nucleases". Current Opinion in Virology. 12: 85–90. doi:10.1016/j.coviro.2015.03.011. PMC 4470805. PMID 25914022.
- ↑ Maggio I, Gonçalves MA (May 2015). "Genome editing at the crossroads of delivery, specificity, and fidelity". Trends in Biotechnology. 33 (5): 280–291. doi:10.1016/j.tibtech.2015.02.011. PMID 25819765.
- ↑ Rath D, Amlinger L, Rath A, Lundgren M (October 2015). "The CRISPR-Cas immune system: biology, mechanisms and applications". Biochimie. 117: 119–128. doi:10.1016/j.biochi.2015.03.025. PMID 25868999.
- ↑ "What Is CRISPR? How Does It Work? Is It Gene Editing?". LiveScience.Tech. 2018-04-30. Archived from the original on 2020-02-06. Retrieved 2020-02-06.
- ↑ 152.0 152.1 Freedman BS, Brooks CR, Lam AQ, Fu H, Morizane R, Agrawal V, et al. (October 2015). "Modelling kidney disease with CRISPR-mutant kidney organoids derived from human pluripotent epiblast spheroids". Nature Communications. 6: 8715. Bibcode:2015NatCo...6.8715F. doi:10.1038/ncomms9715. PMC 4620584. PMID 26493500.
- ↑ Cruz NM, Song X, Czerniecki SM, Gulieva RE, Churchill AJ, Kim YK, et al. (November 2017). "Organoid cystogenesis reveals a critical role of microenvironment in human polycystic kidney disease". Nature Materials. 16 (11): 1112–1119. Bibcode:2017NatMa..16.1112C. doi:10.1038/nmat4994. PMC 5936694. PMID 28967916.
- ↑ Kim YK, Refaeli I, Brooks CR, Jing P, Gulieva RE, Hughes MR, et al. (December 2017). "Gene-Edited Human Kidney Organoids Reveal Mechanisms of Disease in Podocyte Development". Stem Cells. 35 (12): 2366–2378. doi:10.1002/stem.2707. PMC 5742857. PMID 28905451.
- ↑ Bellin M, Casini S, Davis RP, D'Aniello C, Haas J, Ward-van Oostwaard D, et al. (December 2013). "Isogenic human pluripotent stem cell pairs reveal the role of a KCNH2 mutation in long-QT syndrome". The EMBO Journal. 32 (24): 3161–3175. doi:10.1038/emboj.2013.240. PMC 3981141. PMID 24213244.
- ↑ Cai L, Fisher AL, Huang H, Xie Z (December 2016). "CRISPR-mediated genome editing and human diseases". Genes & Diseases. 3 (4): 244–251. doi:10.1016/j.gendis.2016.07.003. PMC 6150104. PMID 30258895.
- ↑ "Seven Diseases That CRISPR Technology Could Cure". Labiotech.eu. 2018-06-25. Archived from the original on 2018-08-21. Retrieved 2018-08-22.
- ↑ 158.0 158.1 "CRISPR/Cas9 and Cancer". Immuno-Oncology News. 2018-04-27. Archived from the original on 2019-02-18. Retrieved 2019-02-18.
- ↑ Crossley M (February 2021). "New CRISPR technology could revolutionise gene therapy, offering new hope to people with genetic diseases". The Conversation. Archived from the original on 2021-02-03. Retrieved 2021-02-03.
- ↑ Cromer MK, Camarena J, Martin RM, Lesch BJ, Vakulskas CA, Bode NM, et al. (April 2021). "Gene replacement of α-globin with β-globin restores hemoglobin balance in β-thalassemia-derived hematopoietic stem and progenitor cells". Nature Medicine. 27 (4): 677–687. doi:10.1038/s41591-021-01284-y. PMC 8265212. PMID 33737751.
- ↑ Xie F, Ye L, Chang JC, Beyer AI, Wang J, Muench MO, Kan YW (September 2014). "Seamless gene correction of β-thalassemia mutations in patient-specific iPSCs using CRISPR/Cas9 and piggyBac". Genome Research. 24 (9): 1526–1533. doi:10.1101/gr.173427.114. PMC 4158758. PMID 25096406.
- ↑ 162.0 162.1 Frangoul H, Altshuler D, Cappellini MD, Chen YS, Domm J, Eustace BK, et al. (January 2021). "CRISPR-Cas9 Gene Editing for Sickle Cell Disease and β-Thalassemia". The New England Journal of Medicine. 384 (3): 252–260. doi:10.1056/NEJMoa2031054. PMID 33283989. S2CID 227521558.
- ↑ Dever DP, Bak RO, Reinisch A, Camarena J, Washington G, Nicolas CE, et al. (November 2016). "CRISPR/Cas9 β-globin gene targeting in human haematopoietic stem cells". Nature. 539 (7629): 384–389. Bibcode:2016Natur.539..384D. doi:10.1038/nature20134. PMC 5898607. PMID 27820943.
- ↑ "CRISPR 'One Shot' Cell Therapy for Hemophilia Developed". GEN. 2018-05-02. Archived from the original on 2018-08-22. Retrieved 2018-08-22.
- ↑ Marangi M, Pistritto G (2018-04-20). "Innovative Therapeutic Strategies for Cystic Fibrosis: Moving Forward to CRISPR Technique". Frontiers in Pharmacology. 9: 396. doi:10.3389/fphar.2018.00396. PMC 5920621. PMID 29731717.
- ↑ Bengtsson NE, Hall JK, Odom GL, Phelps MP, Andrus CR, Hawkins RD, et al. (February 2017). "Muscle-specific CRISPR/Cas9 dystrophin gene editing ameliorates pathophysiology in a mouse model for Duchenne muscular dystrophy". Nature Communications. 8: 14454. Bibcode:2017NatCo...814454B. doi:10.1038/ncomms14454. PMC 5316861. PMID 28195574.
- ↑ Eisenstein M (May 2018). "CRISPR takes on Huntington's disease". Nature. 557 (7707): S42–S43. Bibcode:2018Natur.557S..42E. doi:10.1038/d41586-018-05177-y. PMID 29844549.
- ↑ Dabrowska M, Juzwa W, Krzyzosiak WJ, Olejniczak M (2018). "Precise Excision of the CAG Tract from the Huntingtin Gene by Cas9 Nickases". Frontiers in Neuroscience. 12: 75. doi:10.3389/fnins.2018.00075. PMC 5834764. PMID 29535594.
- ↑ King A (March 2018). "A CRISPR edit for heart disease". Nature. 555 (7695): S23–S25. Bibcode:2018Natur.555.....K. doi:10.1038/d41586-018-02482-4. PMID 29517035.
- ↑ Scudellari M (July 2019). "Self-destructing mosquitoes and sterilized rodents: the promise of gene drives". Nature. 571 (7764): 160–162. Bibcode:2019Natur.571..160S. doi:10.1038/d41586-019-02087-5. PMID 31289403.
- ↑ Abrahimi P, Chang WG, Kluger MS, Qyang Y, Tellides G, Saltzman WM, Pober JS (July 2015). "Efficient gene disruption in cultured primary human endothelial cells by CRISPR/Cas9". Circulation Research. 117 (2): 121–128. doi:10.1161/CIRCRESAHA.117.306290. PMC 4490936. PMID 25940550.
- ↑ "A Year In, 1st Patient To Get Gene Editing For Sickle Cell Disease Is Thriving". NPR.org. Archived from the original on 2020-06-30. Retrieved 2021-02-03.
- ↑ "CRISPR technology to cure sickle cell disease". ScienceDaily. Archived from the original on 2021-01-29. Retrieved 2021-02-03.
- ↑ Uddin F, Rudin CM, Sen T (August 2020). "CRISPR Gene Therapy: Applications, Limitations, and Implications for the Future". Frontiers in Oncology. 10: 1387. doi:10.3389/fonc.2020.01387. PMC 7427626. PMID 32850447.
- ↑ 175.0 175.1 175.2 175.3 "CRISPR Clinical Trials: A 2022 Update". Innovative Genomics Institute (IGI). Archived from the original on 2022-05-18. Retrieved 2022-05-02.
- ↑ "Editas Hits Pause on LCA10 Program in Search of Partner". Archived from the original on 2023-03-27. Retrieved 2023-11-21.
- ↑ Jensen TI, Axelgaard E, Bak RO (June 2019). "Therapeutic gene editing in haematological disorders with CRISPR/Cas9". British Journal of Haematology. 185 (5): 821–835. doi:10.1111/bjh.15851. PMID 30864164. S2CID 76663873.
- ↑ Reardon S (2016). "First CRISPR clinical trial gets green light from US panel". Nature. doi:10.1038/nature.2016.20137. S2CID 89466280.
- ↑ Rosenblum D, Gutkin A, Kedmi R, Ramishetti S, Veiga N, Jacobi AM, et al. (November 2020). "CRISPR-Cas9 genome editing using targeted lipid nanoparticles for cancer therapy". Science Advances. 6 (47): eabc9450. Bibcode:2020SciA....6.9450R. doi:10.1126/sciadv.abc9450. PMC 7673804. PMID 33208369.
- ↑ "Base editing: Revolutionary therapy clears girl's incurable cancer". BBC News. 11 December 2022. Archived from the original on 11 December 2022. Retrieved 11 December 2022.
- ↑ "'Designer cells' reverse one-year-old's cancer". BBC News. 2015-11-05. Archived from the original on 2022-12-11. Retrieved 2022-12-11.
- ↑ "CRISPR Therapeutics and ViaCyte, Inc. Announce First Patient Dosed in…". CRISPR. Archived from the original on 2022-04-30. Retrieved 2022-05-02.
- ↑ Gomaa AA, Klumpe HE, Luo ML, Selle K, Barrangou R, Beisel CL (January 2014). "Programmable removal of bacterial strains by use of genome-targeting CRISPR-Cas systems". mBio. 5 (1): e00928–e00913. doi:10.1128/mBio.00928-13. PMC 3903277. PMID 24473129.
- ↑ Citorik RJ, Mimee M, Lu TK (November 2014). "Sequence-specific antimicrobials using efficiently delivered RNA-guided nucleases". Nature Biotechnology. 32 (11): 1141–1145. doi:10.1038/nbt.3011. hdl:1721.1/100834. PMC 4237163. PMID 25240928.
- ↑ 185.0 185.1 Gholizadeh P, Aghazadeh M, Asgharzadeh M, Kafil HS (November 2017). "Suppressing the CRISPR/Cas adaptive immune system in bacterial infections". European Journal of Clinical Microbiology & Infectious Diseases. 36 (11): 2043–2051. doi:10.1007/s10096-017-3036-2. PMID 28601970. S2CID 22716314.
- ↑ Gibney E (January 2018). "What to expect in 2018: science in the new year". Nature. 553 (7686): 12–13. Bibcode:2018Natur.553...12G. doi:10.1038/d41586-018-00009-5. PMID 29300040.
- ↑ Taylor P (Jan 3, 2019). "J&J takes stake in Locus' CRISPR-based 'Pac-Man' antimicrobials". Fierce Biotech. Archived from the original on 6 March 2019. Retrieved 27 February 2019.
- ↑ Reardon S (June 2017). "Modified viruses deliver death to antibiotic-resistant bacteria". Nature. 546 (7660): 586–587. Bibcode:2017Natur.546..586R. doi:10.1038/nature.2017.22173. PMID 28661508.
- ↑ van Diemen FR, Kruse EM, Hooykaas MJ, Bruggeling CE, Schürch AC, van Ham PM, et al. (June 2016). "CRISPR/Cas9-Mediated Genome Editing of Herpesviruses Limits Productive and Latent Infections". PLOS Pathogens. 12 (6): e1005701. doi:10.1371/journal.ppat.1005701. PMC 4928872. PMID 27362483.
- "Using CRISPR to combat viral infections: a new way to treat herpes?". PLOS Media. August 4, 2016. Archived from the original on May 18, 2023. Retrieved November 21, 2023 – via YouTube.
- ↑ National Institute on Drug Abuse (2020-02-14). "Antiretroviral Therapy Combined With CRISPR Gene Editing Can Eliminate HIV Infection in Mice". National Institute on Drug Abuse. Archived from the original on 2022-03-15. Retrieved 2022-04-18.
- ↑ "First Clinical Trial of CRISPR-Based HIV Therapy Founded on Breakthrough Research at Lewis Katz School of Medicine | Lewis Katz School of Medicine at Temple University". medicine.temple.edu. Archived from the original on 2022-04-12. Retrieved 2022-04-18.
- ↑ Excision BioTherapeutics (2022-03-24). "A Phase 1/2a, Sequential Cohort, Single Ascending Dose Study of the Safety, Tolerability, Biodistribution, and Pharmacodynamics of EBT 101 in Aviremic HIV-1 Infected Adults on Stable Antiretroviral Therapy". Archived from the original on 2023-04-07. Retrieved 2023-11-21.
- ↑ 193.0 193.1 193.2 Science News Staff (December 17, 2015). "And Science's Breakthrough of the Year is ..." news.sciencemag.org. Archived from the original on 2015-12-21. Retrieved 2015-12-21.
- ↑ Mullin E. "Using CRISPR on pigs could make their organs safer for human transplant". MIT Technology Review. Archived from the original on 2017-08-13. Retrieved 2017-09-09.
- ↑ Pinson, Anneline; Xing, Lei; Namba, Takashi; Kalebic, Nereo; Peters, Jula; Oegema, Christina Eugster; Traikov, Sofia; Reppe, Katrin; Riesenberg, Stephan; Maricic, Tomislav; Derihaci, Razvan; Wimberger, Pauline; Pääbo, Svante; Huttner, Wieland B. (2022-09-09). "Human TKTL1 implies greater neurogenesis in frontal neocortex of modern humans than Neanderthals". Science. 377 (6611): eabl6422. doi:10.1126/science.abl6422. ISSN 0036-8075. PMID 36074851. S2CID 252161562. Archived from the original on 2022-11-02. Retrieved 2023-11-21.
- ↑ Mora-Bermúdez, Felipe; Kanis, Philipp; Macak, Dominik; Peters, Jula; Naumann, Ronald; Xing, Lei; Sarov, Mihail; Winkler, Sylke; Oegema, Christina Eugster; Haffner, Christiane; Wimberger, Pauline; Riesenberg, Stephan; Maricic, Tomislav; Huttner, Wieland B.; Pääbo, Svante (2022-07-29). "Longer metaphase and fewer chromosome segregation errors in modern human than Neanderthal brain development". Science Advances. 8 (30): eabn7702. Bibcode:2022SciA....8N7702M. doi:10.1126/sciadv.abn7702. ISSN 2375-2548. PMC 9337762. PMID 35905187.
- ↑ "Scientists discover how humans develop larger brains than other apes". phys.org. Archived from the original on 19 April 2021. Retrieved 19 April 2021.
- ↑ Benito-Kwiecinski S, Giandomenico SL, Sutcliffe M, Riis ES, Freire-Pritchett P, Kelava I, et al. (April 2021). "An early cell shape transition drives evolutionary expansion of the human forebrain". Cell. 184 (8): 2084–2102.e19. doi:10.1016/j.cell.2021.02.050. PMC 8054913. PMID 33765444.
 Available under CC BY 4.0 Archived 2017-10-16 at the Wayback Machine.
Available under CC BY 4.0 Archived 2017-10-16 at the Wayback Machine.
- ↑ Sawal I. "Mini brains genetically altered with CRISPR to be Neanderthal-like". New Scientist. Archived from the original on 10 March 2021. Retrieved 7 March 2021.
- ↑ Trujillo CA, Rice ES, Schaefer NK, Chaim IA, Wheeler EC, Madrigal AA, et al. (February 2021). "Reintroduction of the archaic variant of NOVA1 in cortical organoids alters neurodevelopment". Science. 371 (6530). doi:10.1126/science.aax2537. PMC 8006534. PMID 33574182.
- ↑ Maricic, Tomislav; Helmbrecht, Nelly; Riesenberg, Stephan; Macak, Dominik; Kanis, Philipp; Lackner, Martin; Pugach-Matveeva, Agrafena Daria; Pääbo, Svante (2021-10-15). "Comment on "Reintroduction of the archaic variant of NOVA1 in cortical organoids alters neurodevelopment"". Science. 374 (6565): eabi6060. doi:10.1126/science.abi6060. ISSN 0036-8075. PMID 34648345. S2CID 238990790. Archived from the original on 2023-09-18. Retrieved 2023-11-21.
- ↑ Herai, Roberto H.; Szeto, Ryan A.; Trujillo, Cleber A.; Muotri, Alysson R. (2021-10-15). "Response to Comment on "Reintroduction of the archaic variant of NOVA1 in cortical organoids alters neurodevelopment"". Science. 374 (6565): eabi9881. doi:10.1126/science.abi9881. ISSN 0036-8075. PMID 34648331. S2CID 238990560. Archived from the original on 2023-09-18. Retrieved 2023-11-21.
- ↑ Riesenberg, Stephan; Kanis, Philipp; Macak, Dominik; Wollny, Damian; Düsterhöft, Dorothee; Kowalewski, Johannes; Helmbrecht, Nelly; Maricic, Tomislav; Pääbo, Svante (2023-07-20). "Efficient high-precision homology-directed repair-dependent genome editing by HDRobust". Nature Methods. 20 (9): 1388–1399. doi:10.1038/s41592-023-01949-1. ISSN 1548-7091. PMC 10482697. PMID 37474806.
{{cite journal}}: Check|pmc=value (help) - ↑ Dominguez AA, Lim WA, Qi LS (January 2016). "Beyond editing: repurposing CRISPR-Cas9 for precision genome regulation and interrogation". Nature Reviews. Molecular Cell Biology. 17 (1): 5–15. doi:10.1038/nrm.2015.2. PMC 4922510. PMID 26670017.
- ↑ Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelson T, et al. (January 2014). "Genome-scale CRISPR-Cas9 knockout screening in human cells". Science. 343 (6166): 84–87. Bibcode:2014Sci...343...84S. doi:10.1126/science.1247005. PMC 4089965. PMID 24336571.
- ↑ 206.0 206.1 Pennisi E (August 2013). "The CRISPR craze". News Focus. Science. 341 (6148): 833–836. Bibcode:2013Sci...341..833P. doi:10.1126/science.341.6148.833. PMID 23970676.
- ↑ 207.0 207.1 Zimmer C (2016-06-03). "Scientists Find Form of CRISPR Gene Editing With New Capabilities". The New York Times. ISSN 0362-4331. Archived from the original on 2022-10-25. Retrieved 2016-06-10.
- ↑ Pickar-Oliver A, Gersbach CA (August 2019). "The next generation of CRISPR-Cas technologies and applications". Nature Reviews. Molecular Cell Biology. 20 (8): 490–507. doi:10.1038/s41580-019-0131-5. PMC 7079207. PMID 31147612.
- ↑ Basak J, Nithin C (2015). "Targeting Non-Coding RNAs in Plants with the CRISPR-Cas Technology is a Challenge yet Worth Accepting". Frontiers in Plant Science. 6: 1001. doi:10.3389/fpls.2015.01001. PMC 4652605. PMID 26635829.
- ↑ Akbari OS, Bellen HJ, Bier E, Bullock SL, Burt A, Church GM, et al. (August 2015). "BIOSAFETY. Safeguarding gene drive experiments in the laboratory". Science. 349 (6251): 927–929. Bibcode:2015Sci...349..927A. doi:10.1126/science.aac7932. PMC 4692367. PMID 26229113.
- ↑ Caplan AL, Parent B, Shen M, Plunkett C (November 2015). "No time to waste—the ethical challenges created by CRISPR: CRISPR/Cas, being an efficient, simple, and cheap technology to edit the genome of any organism, raises many ethical and regulatory issues beyond the use to manipulate human germ line cells". EMBO Reports. 16 (11): 1421–1426. doi:10.15252/embr.201541337. PMC 4641494. PMID 26450575.
- ↑ Oye KA, Esvelt K, Appleton E, Catteruccia F, Church G, Kuiken T, et al. (August 2014). "Biotechnology. Regulating gene drives". Science. 345 (6197): 626–628. Bibcode:2014Sci...345..626O. doi:10.1126/science.1254287. PMID 25035410.
- ↑ Gu W, Crawford ED, O'Donovan BD, Wilson MR, Chow ED, Retallack H, DeRisi JL (March 2016). "Depletion of Abundant Sequences by Hybridization (DASH): using Cas9 to remove unwanted high-abundance species in sequencing libraries and molecular counting applications". Genome Biology. 17: 41. doi:10.1186/s13059-016-0904-5. PMC 4778327. PMID 26944702.
- ↑ McDonnell S. "New CRISPR-based tool inserts large DNA sequences at desired sites in cells". Massachusetts Institute of Technology via phys.org. Archived from the original on 18 December 2022. Retrieved 18 December 2022.
- ↑ Yarnall MT, Ioannidi EI, Schmitt-Ulms C, Krajeski RN, Lim J, Villiger L, et al. (November 2022). "Drag-and-drop genome insertion of large sequences without double-strand DNA cleavage using CRISPR-directed integrases". Nature Biotechnology. 41 (4): 500–512. bioRxiv 10.1101/2021.11.01.466786. doi:10.1038/s41587-022-01527-4. PMC 10257351. PMID 36424489. S2CID 253879386.
{{cite journal}}: Check|pmc=value (help) - ↑ Anzalone AV, Randolph PB, Davis JR, Sousa AA, Koblan LW, Levy JM, et al. (December 2019). "Search-and-replace genome editing without double-strand breaks or donor DNA". Nature. 576 (7785): 149–157. Bibcode:2019Natur.576..149A. doi:10.1038/s41586-019-1711-4. PMC 6907074. PMID 31634902.
- ↑ Thulin L (21 October 2019). "A New Gene Editing Tool Could Make CRISPR More Precise". The Smithsonian Magazine. Archived from the original on 30 May 2023. Retrieved 21 November 2023.
- ↑ 218.0 218.1 Cohen J (21 October 2019). "New 'prime' genome editor could surpass CRISPR". Science Magazine. Archived from the original on 20 November 2023. Retrieved 21 November 2023.
- ↑ 219.0 219.1 Yasinski E (21 October 2019). "New "Prime Editing" Method Makes Only Single-Stranded DNA Cuts". The Scientist. Archived from the original on 23 October 2019. Retrieved 21 November 2023.
- ↑ 220.0 220.1 Gallagher J (21 October 2019). "Prime editing: DNA tool could correct 89% of genetic defects". BBC News. Archived from the original on 21 October 2019. Retrieved 21 November 2023.
- ↑ Regalado A (March 5, 2015). "Engineering the Perfect Baby". MIT Technology Review. Archived from the original on March 13, 2015.
- ↑ Baltimore D, Berg P, Botchan M, Carroll D, Charo RA, Church G, et al. (April 2015). "Biotechnology. A prudent path forward for genomic engineering and germline gene modification". Science. 348 (6230): 36–38. Bibcode:2015Sci...348...36B. doi:10.1126/science.aab1028. PMC 4394183. PMID 25791083.
- ↑ Lanphier E, Urnov F, Haecker SE, Werner M, Smolenski J (March 2015). "Don't edit the human germ line". Nature. 519 (7544): 410–411. Bibcode:2015Natur.519..410L. doi:10.1038/519410a. PMID 25810189.
- ↑ Wade N (19 March 2015). "Scientists Seek Ban on Method of Editing the Human Genome". The New York Times. Archived from the original on 19 March 2015. Retrieved 20 March 2015.
The biologists writing in Science support continuing laboratory research with the technique, and few if any scientists believe it is ready for clinical use.
- ↑ Liang P, Xu Y, Zhang X, Ding C, Huang R, Zhang Z, et al. (May 2015). "CRISPR/Cas9-mediated gene editing in human tripronuclear zygotes". Protein & Cell. 6 (5): 363–372. doi:10.1007/s13238-015-0153-5. PMC 4417674. PMID 25894090.
- ↑ Kolata G (23 April 2015). "Chinese Scientists Edit Genes of Human Embryos, Raising Concerns". The New York Times. Archived from the original on 24 April 2015. Retrieved 24 April 2015.
- ↑ 227.0 227.1 Cyranoski D, Reardon S (2015). "Chinese scientists genetically modify human embryos". Nature. doi:10.1038/nature.2015.17378. S2CID 87604469.
- ↑ Regalado A (2016-05-08). "Chinese Researchers Experiment with Making HIV-Proof Embryos". MIT Technology Review. Archived from the original on 2016-04-20. Retrieved 2016-06-10.
- ↑ "International Summit on Gene Editing". National Academies of Sciences, Engineering, and Medicine. 3 December 2015. Archived from the original on 5 December 2015. Retrieved 3 December 2015.
- ↑ Brokowski C (April 2018). "Do CRISPR Germline Ethics Statements Cut It?". The CRISPR Journal. 1 (2): 115–125. doi:10.1089/crispr.2017.0024. PMC 6694771. PMID 31021208.
- ↑ Begley S (28 November 2018). "Amid uproar, Chinese scientist defends creating gene-edited babies". STAT. Archived from the original on 31 January 2019. Retrieved 21 November 2023.
- ↑ Sample I (13 March 2019). "Scientists call for global moratorium on gene editing of embryos". Theguardian.com. Archived from the original on 14 March 2019. Retrieved 14 March 2019.
- ↑ Callaway E (February 2016). "UK scientists gain licence to edit genes in human embryos". Nature. 530 (7588): 18. Bibcode:2016Natur.530...18C. doi:10.1038/nature.2016.19270. PMID 26842037.
- ↑ McHughen A, Smyth S (January 2008). "US regulatory system for genetically modified [genetically modified organism (GMO), rDNA or transgenic] crop cultivars". Plant Biotechnology Journal. 6 (1): 2–12. doi:10.1111/j.1467-7652.2007.00300.x. PMID 17956539. S2CID 3210837.
- ↑ USDA. "Re: Request to confirm" (PDF). Archived (PDF) from the original on 2023-06-11. Retrieved 2023-11-21.
- ↑ Waltz E (April 2016). "Gene-edited CRISPR mushroom escapes US regulation". Nature. 532 (7599): 293. Bibcode:2016Natur.532..293W. doi:10.1038/nature.2016.19754. PMID 27111611.
- ↑ Ledford H (April 2016). "Gene-editing surges as US rethinks regulations". Nature. 532 (7598): 158–159. Bibcode:2016Natur.532..158L. doi:10.1038/532158a. PMID 27075074.
- ↑ Brown KV (1 February 2017). "The FDA Is Cracking Down On Rogue Genetic Engineers". Gizmodo. Archived from the original on 5 February 2017. Retrieved 5 February 2017.
- ↑ "Guidance for Industry #187 / Regulation of Intentionally Altered Genomic DNA in Animals" (PDF). Food and Drug Administration. 2020-02-11. Archived (PDF) from the original on 2019-01-08. Retrieved 2023-11-21.
- ↑ Cyranoski D (August 2017). "China's embrace of embryo selection raises thorny questions". Nature. 548 (7667): 272–274. Bibcode:2017Natur.548..272C. doi:10.1038/548272a. PMID 28816265.
- ↑ Peng Y (June 2016). "The morality and ethics governing CRISPR-Cas9 patents in China". Nature Biotechnology. 34 (6): 616–618. doi:10.1038/nbt.3590. PMID 27281418. S2CID 38509820.
- ↑ Rana P, Marcus A, Fan W (2018-01-21). "China, Unhampered by Rules, Races Ahead in Gene-Editing Trials". Wall Street Journal. ISSN 0099-9660. Archived from the original on 2018-04-07. Retrieved 2018-01-23.
- ↑ Talbot D (2016). "Precise Gene Editing in Plants/ 10 Breakthrough Technologies 2016". MIT Technology review. Massachusetts Institute of Technology. Archived from the original on November 5, 2017. Retrieved 18 March 2016.
- ↑ Larson C, Schaffer A (2014). "Genome Editing/ 10 Breakthrough Technologies 2014". Massachusetts Institute of Technology. Archived from the original on December 5, 2016. Retrieved 18 March 2016.
- ↑ "Tang Prize Laureates". www.tang-prize.org. Archived from the original on 2018-08-06. Retrieved 2018-08-05.
- ↑ "Press release: The Nobel Prize in Chemistry 2020". Nobel Foundation. Archived from the original on 15 January 2021. Retrieved 7 October 2020.
- CS1 errors: PMC
- CS1: long volume value
- Webarchive template wayback links
- Articles containing potentially dated statements from November 2013
- Articles with invalid date parameter in template
- All articles containing potentially dated statements
- Articles containing potentially dated statements from 2015
- Articles containing potentially dated statements from December 2014
- Articles containing potentially dated statements from August 2017
- Articles with hatnote templates targeting a nonexistent page
- Articles containing potentially dated statements from 2017
- Articles containing potentially dated statements from August 2016
- Biotechnology
- Genetic engineering
- Genome editing

